
Room 36-776A
50 Vassar St.
(MIT building 36)
Cambridge, MA 02139
Hi! I am currently a fifth-year Ph.D. student at MIT majoring in Electrical Engineering and Computer Science. My research focuses on machine learning and developing robust and efficient algorithms driven by clinical problems. Applications include motion-robust 3D rendering of the human brain, real-time quality assessment in MR scans as well as pose estimation and motion characterization of fetuses. I am advised by Prof. Elfar Adalsteinsson and collaborate closely with Prof. Polina Golland and Prof. P. Ellen Grant.
I also did summer internships at Google and Meta, working on automated Ads bidding and large-scale video recommendation systems respectively.
Prior to MIT, I received my Bachelor’s degree from Tsinghua University in 2018. I also spent a summer as a research assistant at Stanford, where I was advised by Prof. John Pauly and Prof. Greg Zaharchuk.
news
| Apr 25, 2023 | We just released NeSVoR v0.2.0. Check it out! |
|---|---|
| Apr 19, 2023 | I successfully defended my doctoral thesis today! |
| Mar 16, 2023 | Our paper entitled “Latent Signal Models: Learning Compact Representations of Signal Evolution for Improved Time-Resolved, Multi-contrast MRI” was accepted by MRM! |
| Feb 21, 2023 | Our paper entitled “Zero-Shot Self-Supervised Joint Temporal Image and Sensitivity Map Reconstruction via Linear Latent Space” was accepted by MIDL 2023! |
| Jan 9, 2023 | Our paper entitled “NeSVoR: Implicit Neural Representation for Slice-to-Volume Reconstruction in MRI” was accepted by IEEE TMI! |
selected publications
-

NeSVoR: Implicit Neural Representation for Slice-to-Volume Reconstruction in MRI
IEEE Transactions on Medical Imaging, 2023
Reconstructing 3D MR volumes from multiple motion-corrupted stacks of 2D slices has shown promise in imaging of moving subjects, e.g., fetal MRI. However, existing slice-to-volume reconstruction methods are time-consuming, especially when a high-resolution volume is desired. Moreover, they are still vulnerable to severe subject motion and when image artifacts are present in acquired slices. In this work, we present NeSVoR, a resolution-agnostic slice-to-volume reconstruction method, which models the underlying volume as a continuous function of spatial coordinates with implicit neural representation. To improve robustness to subject motion and other image artifacts, we adopt a continuous and comprehensive slice acquisition model that takes into account rigid inter-slice motion, point spread function, and bias fields. NeSVoR also estimates pixel-wise and slice-wise variances of image noise and enables removal of outliers during reconstruction and visualization of uncertainty. Extensive experiments are performed on both simulated and in vivo data to evaluate the proposed method. Results show that NeSVoR achieves state-of-the-art reconstruction quality while providing two to ten-fold acceleration in reconstruction times over the state-of-the-art algorithms.
-

SVoRT: Iterative Transformer for Slice-to-Volume Registration in Fetal Brain MRI
In Medical Image Computing and Computer Assisted Intervention – MICCAI 2022
Volumetric reconstruction of fetal brains from multiple stacks of MR slices, acquired in the presence of almost unpredictable and often severe subject motion, is a challenging task that is highly sensitive to the initialization of slice-to-volume transformations. We propose a novel slice-to-volume registration method using Transformers trained on synthetically transformed data, which model multiple stacks of MR slices as a sequence. With the attention mechanism, our model automatically detects the relevance between slices and predicts the transformation of one slice using information from other slices. We also estimate the underlying 3D volume to assist slice-to-volume registration and update the volume and transformations alternately to improve accuracy. Results on synthetic data show that our method achieves lower registration error and better reconstruction quality compared with existing state-of-the-art methods. Experiments with real-world MRI data are also performed to demonstrate the ability of the proposed model to improve the quality of 3D reconstruction under severe fetal motion.
-
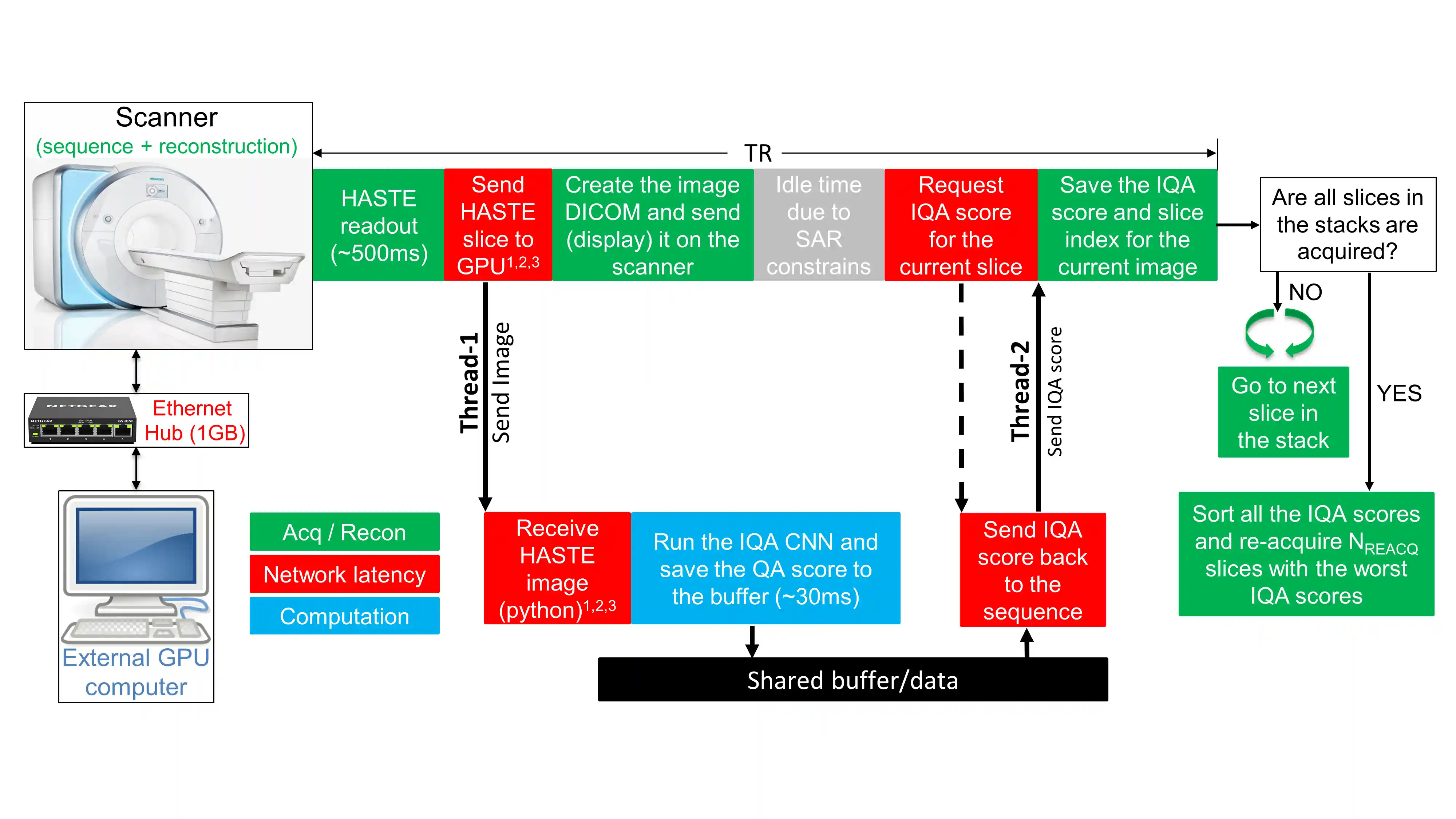
Automated Detection and Reacquisition of Motion-Degraded Images in Fetal HASTE Imaging at 3 T
Borjan Gagoski*, Junshen Xu*, Paul Wighton, M. Dylan Tisdall, and 6 more authors
Magnetic Resonance in Medicine, 2022
Purpose: Fetal brain Magnetic Resonance Imaging suffers from unpredictable and unconstrained fetal motion that causes severe image artifacts even with half-Fourier single-shot fast spin echo (HASTE) readouts. This work presents the implementation of a closed-loop pipeline that automatically detects and reacquires HASTE images that were degraded by fetal motion without any human interaction. Methods: A convolutional neural network that performs automatic image quality assessment (IQA) was run on an external GPU-equipped computer that was connected to the internal network of the MRI scanner. The modified HASTE pulse sequence sent each image to the external computer, where the IQA convolutional neural network evaluated it, and then the IQA score was sent back to the sequence. At the end of the HASTE stack, the IQA scores from all the slices were sorted, and only slices with the lowest scores (corresponding to the slices with worst image quality) were reacquired. Results: The closed-loop HASTE acquisition framework was tested on 10 pregnant mothers, for a total of 73 acquisitions of our modified HASTE sequence. The IQA convolutional neural network, which was successfully employed by our modified sequence in real time, achieved an accuracy of 85.2% and area under the receiver operator characteristic of 0.899. Conclusion: The proposed acquisition/reconstruction pipeline was shown to successfully identify and automatically reacquire only the motion degraded fetal brain HASTE slices in the prescribed stack. This minimizes the overall time spent on HASTE acquisitions by avoiding the need to repeat the entire stack if only few slices in the stack are motion-degraded.
-
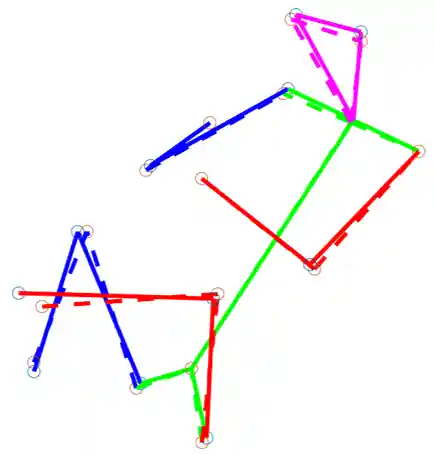
Fetal Pose Estimation in Volumetric MRI using a 3D Convolution Neural Network
In Medical Image Computing and Computer Assisted Intervention – MICCAI 2019
The performance and diagnostic utility of magnetic resonance imaging (MRI) in pregnancy is fundamentally constrained by fetal motion. Motion of the fetus, which is unpredictable and rapid on the scale of conventional imaging times, limits the set of viable acquisition techniques to single-shot imaging with severe compromises in signal-to-noise ratio and diagnostic contrast, and frequently results in unacceptable image quality. Surprisingly little is known about the characteristics of fetal motion during MRI and here we propose and demonstrate methods that exploit a growing repository of MRI observations of the gravid abdomen that are acquired at low spatial resolution but relatively high temporal resolution and over long durations (10–30 min). We estimate fetal pose per frame in MRI volumes of the pregnant abdomen via deep learning algorithms that detect key fetal landmarks. Evaluation of the proposed method shows that our framework achieves quantitatively an average error of 4.47 mm and 96.4% accuracy (with error less than 10 mm). Fetal pose estimation in MRI time series yields novel means of quantifying fetal movements in health and disease, and enables the learning of kinematic models that may enhance prospective mitigation of fetal motion artifacts during MRI acquisition.
-
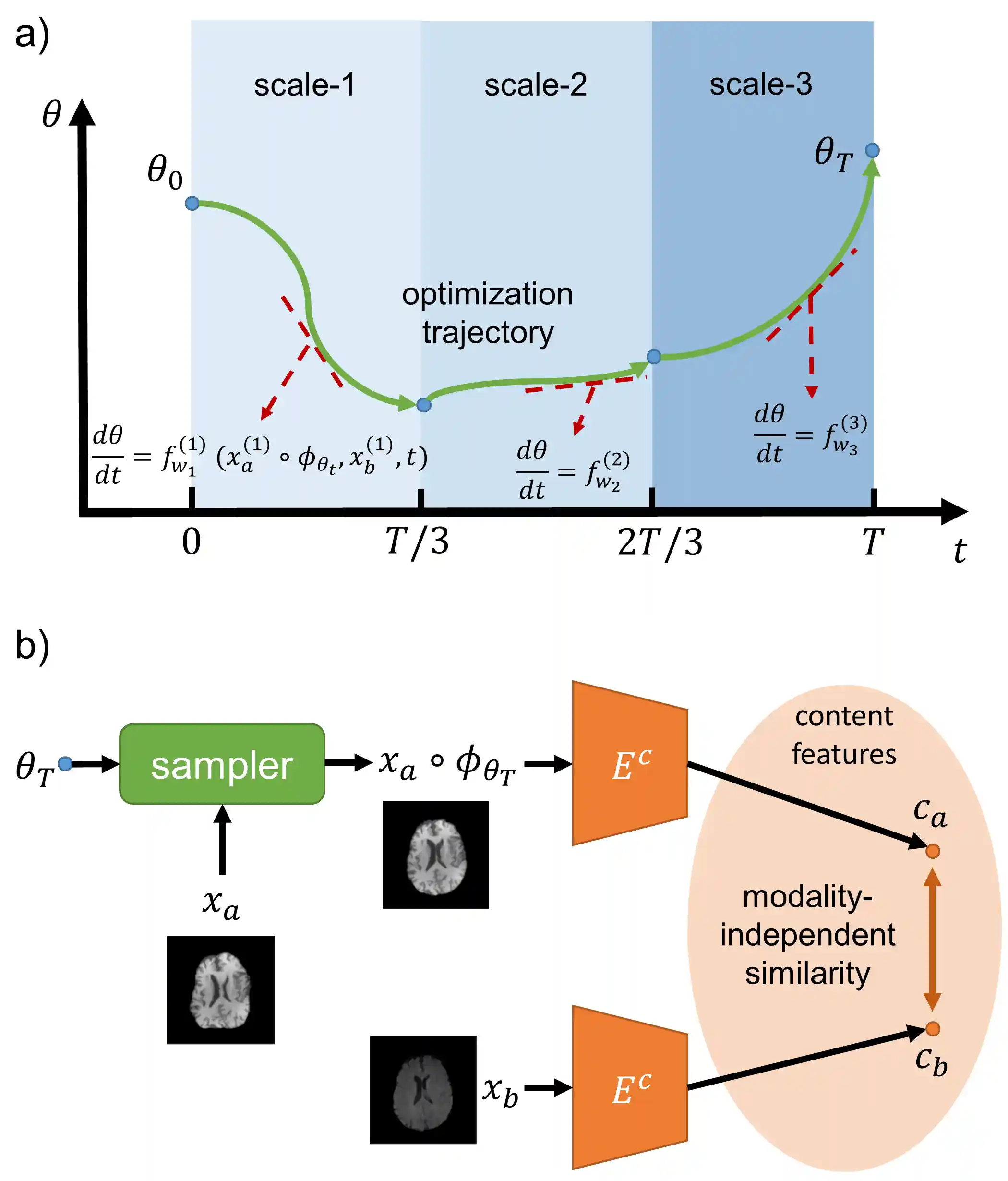
Multi-Scale Neural ODEs for 3D Medical Image Registration
Junshen Xu, Eric Z. Chen, Xiao Chen, Terrence Chen, and 1 more author
In Medical Image Computing and Computer Assisted Intervention – MICCAI 2021
Image registration plays an important role in medical image analysis. Conventional optimization based methods provide an accurate estimation due to the iterative process at the cost of expensive computation. Deep learning methods such as learn-to-map are much faster but either iterative or coarse-to-fine approach is required to improve accuracy for handling large motions. In this work, we proposed to learn a registration optimizer via a multi-scale neural ODE model. The inference consists of iterative gradient updates similar to a conventional gradient descent optimizer but in a much faster way, because the neural ODE learns from the training data to adapt the gradient efficiently at each iteration. Furthermore, we proposed to learn a modal-independent similarity metric to address image appearance variations across different image contrasts. We performed evaluations through extensive experiments in the context of multi-contrast 3D MR images from both public and private data sources and demonstrate the superior performance of our proposed methods.