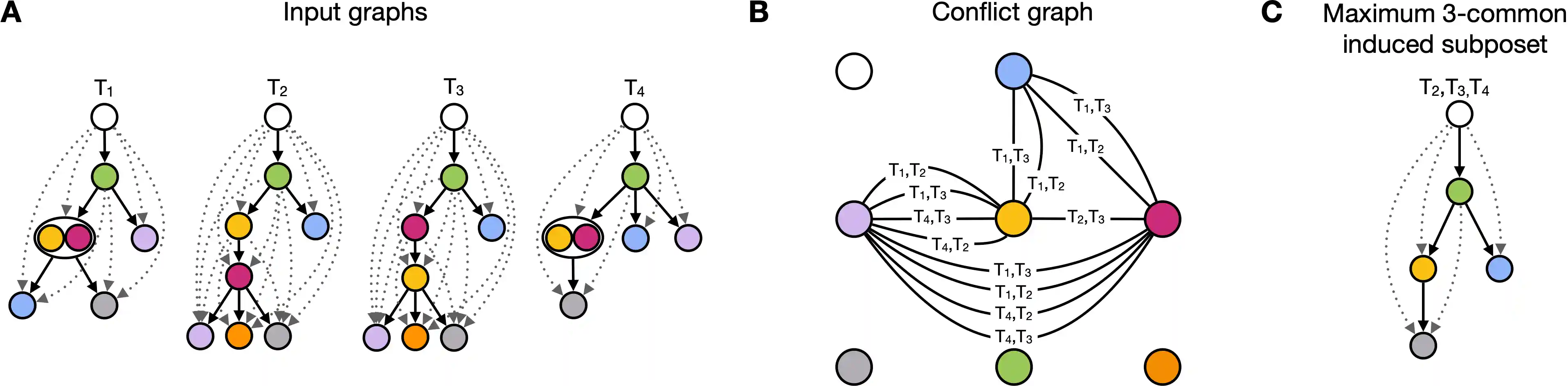
POsets for Temporal Trajectory Resolution (POTTR) is a combinatorial method to identify maximum recurrent trajectories in biological processes described by sequences of events with a temporal ordering. POTTR models these processes, such as tumor formation or cell differentiation, as incomplete posets. A recurrent trajectory is a set of events, like mutations, that follow the same incomplete partial order across distinct incomplete posets.
We model conflicting orders between shared events in a conflict graph. POTTR identifies maximum independent sets in subgraphs of the conflict graph, which directly corresponds to a maximum recurrent trajectory shared in a subset of the input. Since the input data might be very heterogeneous, POTTR searches for recurrent trajectories that are shared in at least $k$ incomplete posets.
Setup
Conda
The easiest way to get POTTR running is by using a conda distribution like miniconda.
After setting up conda (installation + initialization for your shell), create a new environment with the provided environment.yaml which contains the required dependencies for POTTR.
In the code directory execute:
conda env create -f environment.yaml conda activate pottr_env
Gurobi
POTTR uses the Gurobi solver for which a license is required. Further information can be found here.
Data
MASTRO comparison
We provide tumor trees of 89 NSCLC and 120 AML patients in the data/data_mastro directory, which we originally obtained from MASTRO [1].
TRACERx data
The phylogenetic trees from TRACERx are available at zenodo. Download and unpack the zip file from zenodo first. Then follow the instructions in here to create the phylogenetic trees for the execution with POTTR.
Cell differentiation data
We provide cell differentiation maps generated with Carta [2] in the data/data_carta directory in graph exchange XML format, which can be read in by POTTR directly.
Execution
In the code directory, run POTTR via the ./run_POTTR.py Python script.
Program arguments
The following arguments are available:
| Argument | Argument long form | Description |
|---|---|---|
| -h | --help | show help message and exit |
| -o | --output-path | Path to store output files |
| -d | --dags | File or directory containing transitively closed DAGs (incomplete posets) |
| -k | --k | Number k of incomplete posets to search for common trajectory |
| -rt | --resolution_threshold | Optional threshold of orders a ≺ b that must be observed in the data to resolve a hidden order a ~ b to a ≺ b (default=1) |
| -rf | --resolution_frequency | Optional flag to only resolve hidden orders a ~ b in the direction of the most frequent order, i.e. either a ≺ b or b ≺ a, but not both |
| -c | --cores | Number cores / threads Gurobi should use; default 0, Gurobi will use all available cores |
| -parallel | --parallelize | Enable parallel processing for creating conflict graph |
| -pool | --solution-pool-size | Solution pool size for Gurobi to retrieve multiple solutions |
| -v | --verbose | Increase output verbosity |
| -dots | --draw_dots | Create trajectory png files (only recommended for small instances) |
Especially in tumor trees inferred from bulk sequencing data, mutation clusters are very common. Mutations in such a cluster are related by a hidden order, which POTTR can resolve to a certain order.
Generally, it is sufficient for POTTR to observe one tumor tree with a known order for a mutation pair to resolve the hidden relation between this pair in a different tumor graph.
However, to increase confidence in the resolution, you might want to require a larger number of tumor trees supporting a certain order to resolve a hidden order to that order.
Or, in case that a ~ b, a ≺ b, and b ≺ a are observed in the data, you probably wish to resolve a cluster only in the direction of the most frequent order a ≺ b or b ≺ a, but not in both directions.
This you can achieve through the parameters --resolution_threshold and --resolution_frequency.
Example execution with test data
In the code directory execute:
python run_POTTR.py --dags ../Data/test_data/ --output-path ../Data/output -k 3 -v
Program ouput
output
├── converted_graphs.txt -- identified maximum recurrent trajectories and all input DAGs supporting each trajectory
├── number_of_distinct_dags_per_sample.csv -- overview of distinct DAGs per sample read from the input
├── processed_graphs
│ └── processed_graphs_support.csv
├── significance_output.txt -- statistical significance values of maximum recurrent trajectories
└── trajectories_gexf
├── 0_trajectory.gexf -- maximum recurrent trajectory identified by POTTR
└── traj_graphs_names.csv -- matching between trajectories and supporting input DAGs
References
- Leonardo Pellegrina, Fabio Vandin, Discovering significant evolutionary trajectories in cancer phylogenies, Bioinformatics, Volume 38, Issue Supplement_2, September 2022, Pages ii49–ii55, https://doi.org/10.1093/bioinformatics/btac467
- Palash Sashittal, et al., Inferring cell differentiation maps from lineage tracing data, International Conference on Research in Computational Molecular Biology, Cham: Springer Nature Switzerland, 2025, https://doi.org/10.1007/978-3-031-90252-9_29