The Torso Processing ToolBox (TPTBox) is a multi-functional package to handle any sort of bids-conform dataset (CT, MRI, ...) It can find, filter, search any BIDS_Family and subjects, and has many functionalities, among them:
- Easily loop over datasets, and the required files
- Read, Write Niftys, centroid jsons, ...
- Reorient, Resample, Shift Niftys, Centroids, labels
- Modular 2D snapshot generation (different views, MIPs, ...)
- 3D Mesh generation from segmentation and snapshots from them
- Registration
- Logging everything consistently
- ...
Install the package
conda create -n 3.10 python=3.10
conda activate 3.10
pip install TPTBox
# Optional dependency Registration
pip install hf-deepaliInstall via github:
(you should be in the project folder)
pip install poetry poetry install
or: Develop mode is really, really nice:
pip install poetry poetry install --with dev
Functionalities
Each folder in this package represents a different functionality.
The top-level-hierarchy incorporates the most important files, the BIDS_files.
BIDS_Files
This file builds a data model out of the BIDS file names.
It can load a dataset as a BIDS_Global_info file, from which search queries and loops over the dataset can be started.
See tutorial_BIDS_files.ipynb for details.
bids_constants
Defines constants for the BIDS nomenclature (sequence-splitting keys, naming conventions...)
vert_constants
Contains definitions and sort order for our intern labels, for vertebrae, POI, ...
Rotation and Resampling
Example rotate and resample.
from TPTBox import NII nii = NII.load("...path/xyz.nii.gz", seg=True) # R right, L left # S superior/up, I inferior/down # A anterior/front, P posterior/back img_rot = nii.reorient(axcodes_to=("P", "I", "R")) img_scale = nii.rescale((1.5, 5, 1)) # in mm as currently rotated # resample to an other image img_resampled_to_other = nii.resample_from_to(img_scale) nii.get_array() # get numpy array nii.affine # Affine matrix nii.header # NIFTY header nii.orientation # Orientation in 3-Letters nii.zoom # Scale of the three image axis nii.shape #shape
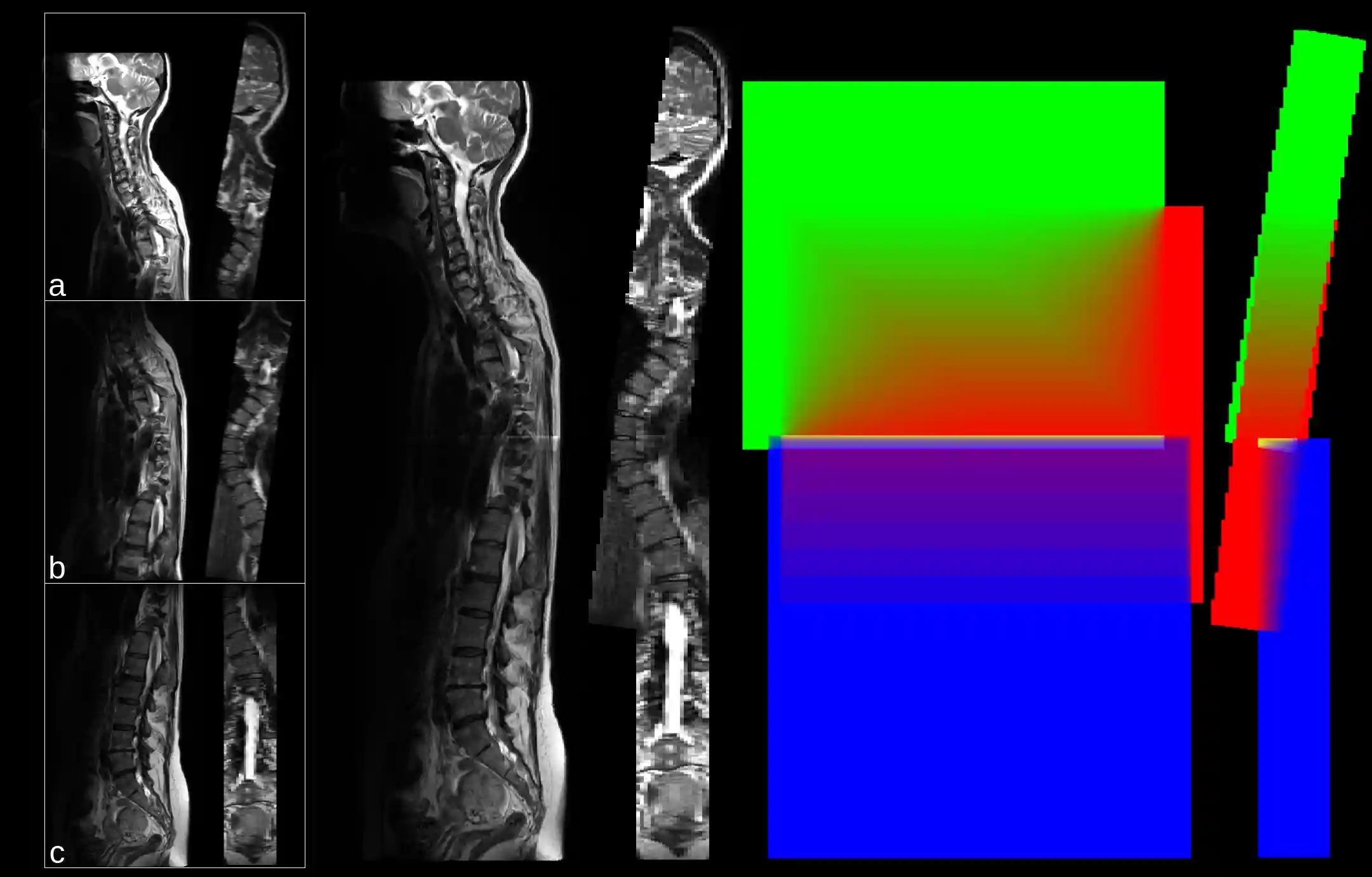
Stitching
Python function and script for arbitrary image stitching. See Details
Spineps and Points of Interests (POI) integration
 For our Spine segmentation pipline follow the installation of SPINEPS.
Image Source: Rule-based Key-Point Extraction for MR-Guided Biomechanical Digital Twins of the Spine;
For our Spine segmentation pipline follow the installation of SPINEPS.
Image Source: Rule-based Key-Point Extraction for MR-Guided Biomechanical Digital Twins of the Spine;
SPINEPS will produce two mask: instance and semantic labels. With these we can compute our POIs. There are either center of mass points or surface points with bioloical meaning. See Validation of a Patient-Specific Musculoskeletal Model for Lumbar Load Estimation Generated by an Automated Pipeline From Whole Body CT
from TPTBox import NII, POI, Location, POI_Global, calc_poi_from_subreg_vert from TPTBox.core.vert_constants import v_name2idx from TPTBox.segmentation.spineps import run_spineps_single # This requires that spineps is installed output_paths = run_spineps_single( "file-path-of_T2w.nii.gz", model_semantic="t2w", ignore_compatibility_issues=True, ) out_spine = output_paths["out_spine"] out_vert = output_paths["out_vert"] semantic_nii = NII.load(out_spine, seg=True) instance_nii = NII.load(out_vert, seg=True) poi = calc_poi_from_subreg_vert( instance_nii, semantic_nii, subreg_id=[ Location.Vertebra_Full, Location.Arcus_Vertebrae, Location.Spinosus_Process, Location.Costal_Process_Left, Location.Costal_Process_Right, Location.Superior_Articular_Left, Location.Superior_Articular_Right, Location.Inferior_Articular_Left, Location.Inferior_Articular_Right, # Location.Vertebra_Corpus_border, CT only Location.Vertebra_Corpus, Location.Vertebra_Disc, Location.Muscle_Inserts_Spinosus_Process, Location.Muscle_Inserts_Transverse_Process_Left, Location.Muscle_Inserts_Transverse_Process_Right, Location.Muscle_Inserts_Vertebral_Body_Left, Location.Muscle_Inserts_Vertebral_Body_Right, Location.Muscle_Inserts_Articulate_Process_Inferior_Left, Location.Muscle_Inserts_Articulate_Process_Inferior_Right, Location.Muscle_Inserts_Articulate_Process_Superior_Left, Location.Muscle_Inserts_Articulate_Process_Superior_Right, Location.Ligament_Attachment_Point_Anterior_Longitudinal_Superior_Median, Location.Ligament_Attachment_Point_Posterior_Longitudinal_Superior_Median, Location.Ligament_Attachment_Point_Anterior_Longitudinal_Inferior_Median, Location.Ligament_Attachment_Point_Posterior_Longitudinal_Inferior_Median, Location.Additional_Vertebral_Body_Middle_Superior_Median, Location.Additional_Vertebral_Body_Posterior_Central_Median, Location.Additional_Vertebral_Body_Middle_Inferior_Median, Location.Additional_Vertebral_Body_Anterior_Central_Median, Location.Ligament_Attachment_Point_Anterior_Longitudinal_Superior_Left, Location.Ligament_Attachment_Point_Posterior_Longitudinal_Superior_Left, Location.Ligament_Attachment_Point_Anterior_Longitudinal_Inferior_Left, Location.Ligament_Attachment_Point_Posterior_Longitudinal_Inferior_Left, Location.Additional_Vertebral_Body_Middle_Superior_Left, Location.Additional_Vertebral_Body_Posterior_Central_Left, Location.Additional_Vertebral_Body_Middle_Inferior_Left, Location.Additional_Vertebral_Body_Anterior_Central_Left, Location.Ligament_Attachment_Point_Anterior_Longitudinal_Superior_Right, Location.Ligament_Attachment_Point_Posterior_Longitudinal_Superior_Right, Location.Ligament_Attachment_Point_Anterior_Longitudinal_Inferior_Right, Location.Ligament_Attachment_Point_Posterior_Longitudinal_Inferior_Right, Location.Additional_Vertebral_Body_Middle_Superior_Right, Location.Additional_Vertebral_Body_Posterior_Central_Right, Location.Additional_Vertebral_Body_Middle_Inferior_Right, Location.Additional_Vertebral_Body_Anterior_Central_Right, Location.Ligament_Attachment_Point_Flava_Superior_Median, Location.Ligament_Attachment_Point_Flava_Inferior_Median, Location.Vertebra_Direction_Posterior, Location.Vertebra_Direction_Inferior, Location.Vertebra_Direction_Right, ], ) poi = poi.round(2) print("Vertebra T4 Vertebra Corpus Center of mass:", poi[v_name2idx["T4"], Location.Vertebra_Corpus]) print("The id number of T4 Vertebra_Corpus is ", v_name2idx["T4"], Location.Vertebra_Corpus.value) # rescale/reorante local poi like nii poi_new = poi.reorient(("P", "I", "R")).rescale((1, 1, 1)) # Local and global POIs can be rescaled to a target spacing with: poi_new = poi.resample_from_to(other_nii_or_poi) # local to global poi global_poi = poi.to_global(itk_coords=True) # You can save global pois in mrk.json format for import and editing in slicer. global_poi.save_mrk("FILE.mrk.json", glyphScale=3.0) # Import as a Markup in slicer; To make points editable you must click on the "lock" symbol under Markups - Control Points - Interaction # Save in our format: poi.save(poi_path) # Loading local/global Poi poi = POI.load(poi_path) poi = POI_Global.load(poi_path)
Snapshot2D Spine
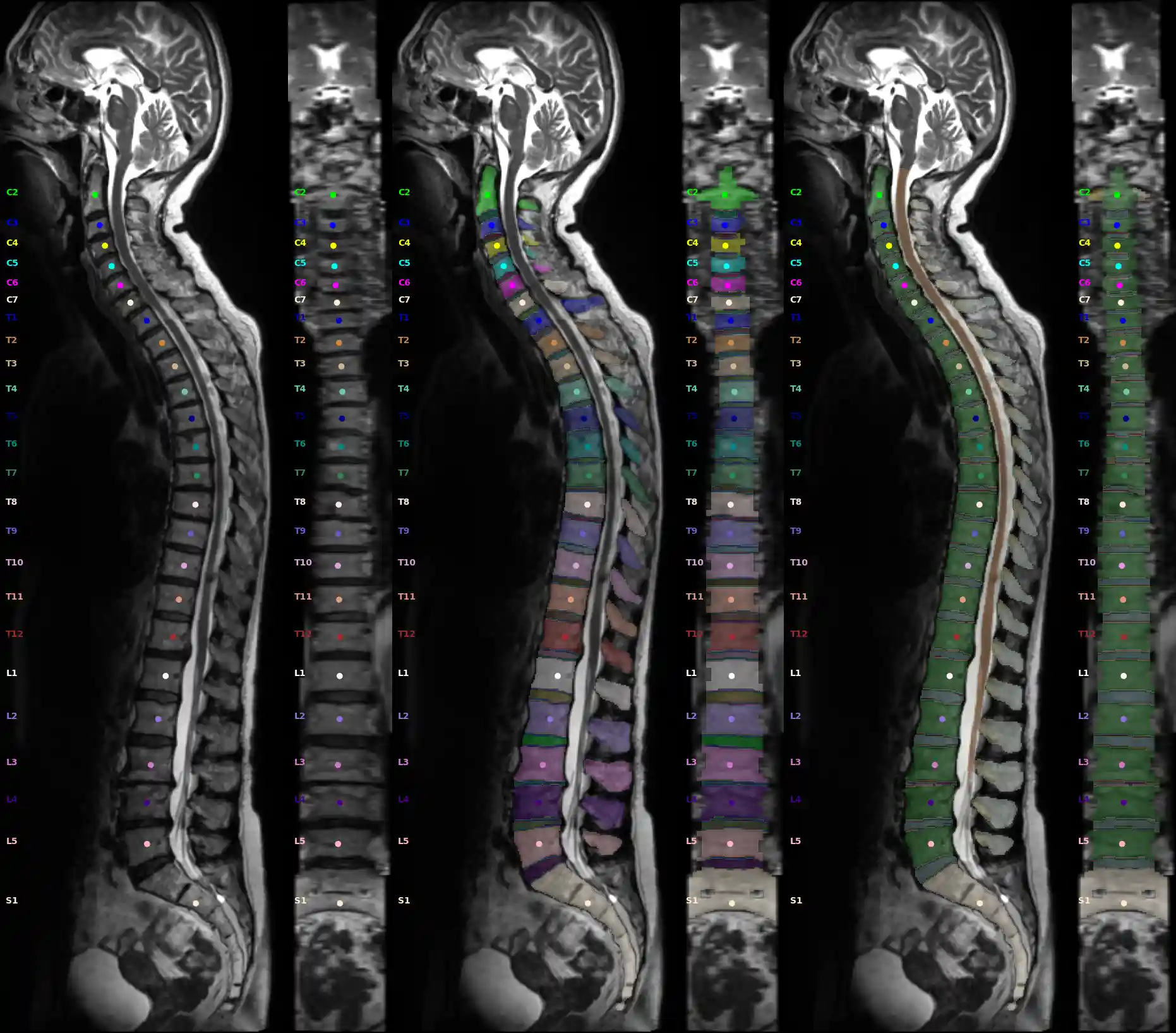 The snapshot function automatically generates sag, cor, axial cuts in the center of a segmentation.
The snapshot function automatically generates sag, cor, axial cuts in the center of a segmentation.
from TPTBox.spine.snapshot2D import Snapshot_Frame, create_snapshot ct = Path("Path to CT") mri = Path("Path to MRI") vert = Path("Path to Vertebra segmentation") subreg = Path("Path to Vertebra subregions") poi_ct = Path("Path to Vertebra poi") poi_mr = Path("Path to Vertebra poi") ct_frame = Snapshot_Frame(image=ct, segmentation=vert, centroids=poi_ct, mode="CT", coronal=True, axial=True) mr_frame = Snapshot_Frame(image=mri, segmentation=vert, centroids=poi_mr, mode="MRI", coronal=True, axial=True) create_snapshot(snp_path="snapshot.jpg", frames=[ct_frame, mr_frame])
Snapshot3D
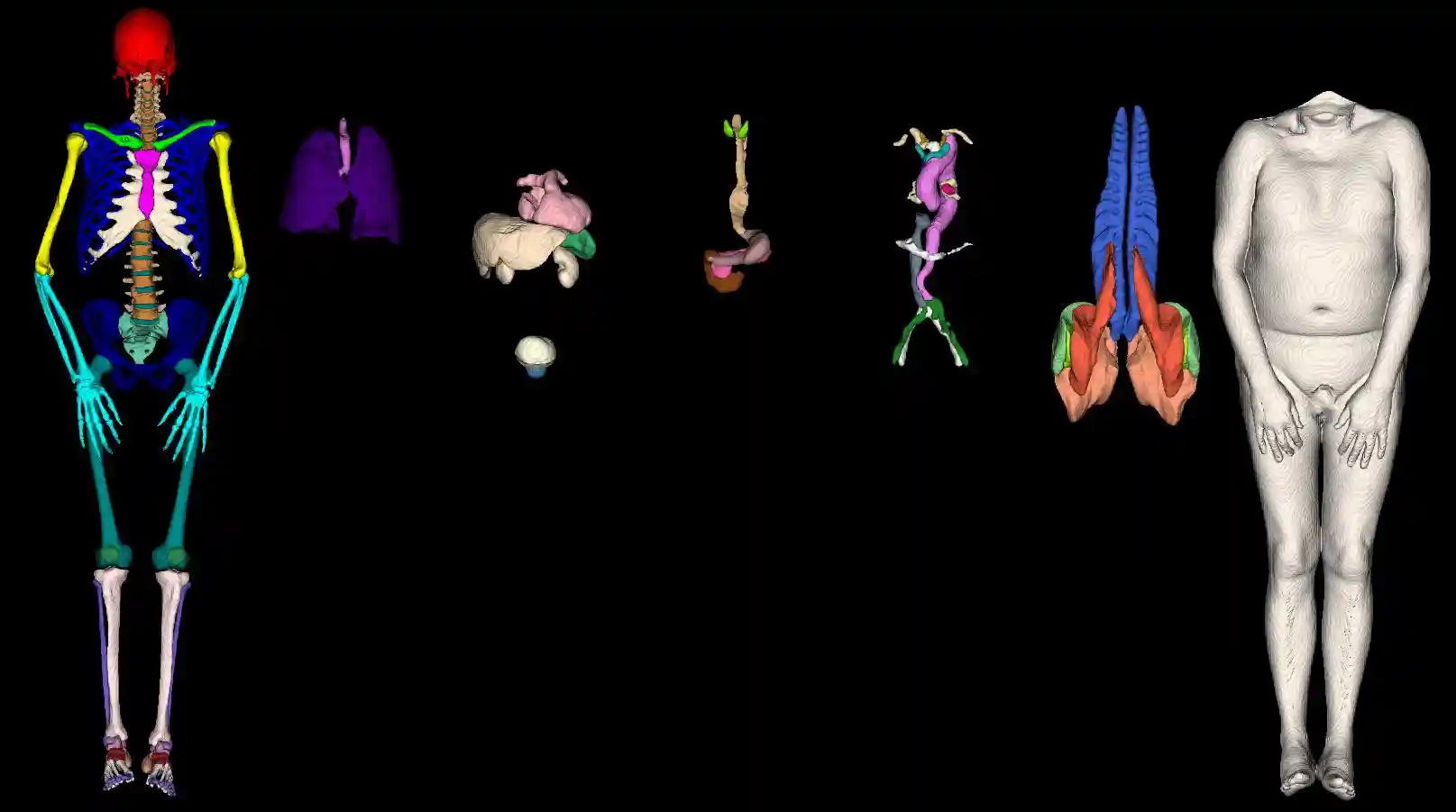 Requires additonal python packages: vtk fury xvfbwrapper
Requires additonal python packages: vtk fury xvfbwrapper
from TPTBox.mesh3D.snapshot3D import make_snapshot3D, make_snapshot3D_parallel # all segmentation; view give the rotation of an image make_snapshot3D("sub-101000_msk.nii.gz", "snapshot3D.png", view=["A", "L", "P", "R"]) # Select witch segmentation per panel are chosen. make_snapshot3D("sub-101000_msk.nii.gz", "snapshot3D_v2.png", view=["A"], ids_list=[[1, 2], [3]]) # we proviede a implementation to process multiple images at the same time. make_snapshot3D_parallel(["a.nii.gz", "b.nii.gz"], ["snp_a.png", "snp_b.png"], view=["A"])