Calculate the similarity between a large number of mass spectra using a GPU. SimMS aims to provide very fast replacements for commonly used similarity functions in matchms.
How SimMS works, in a nutshell
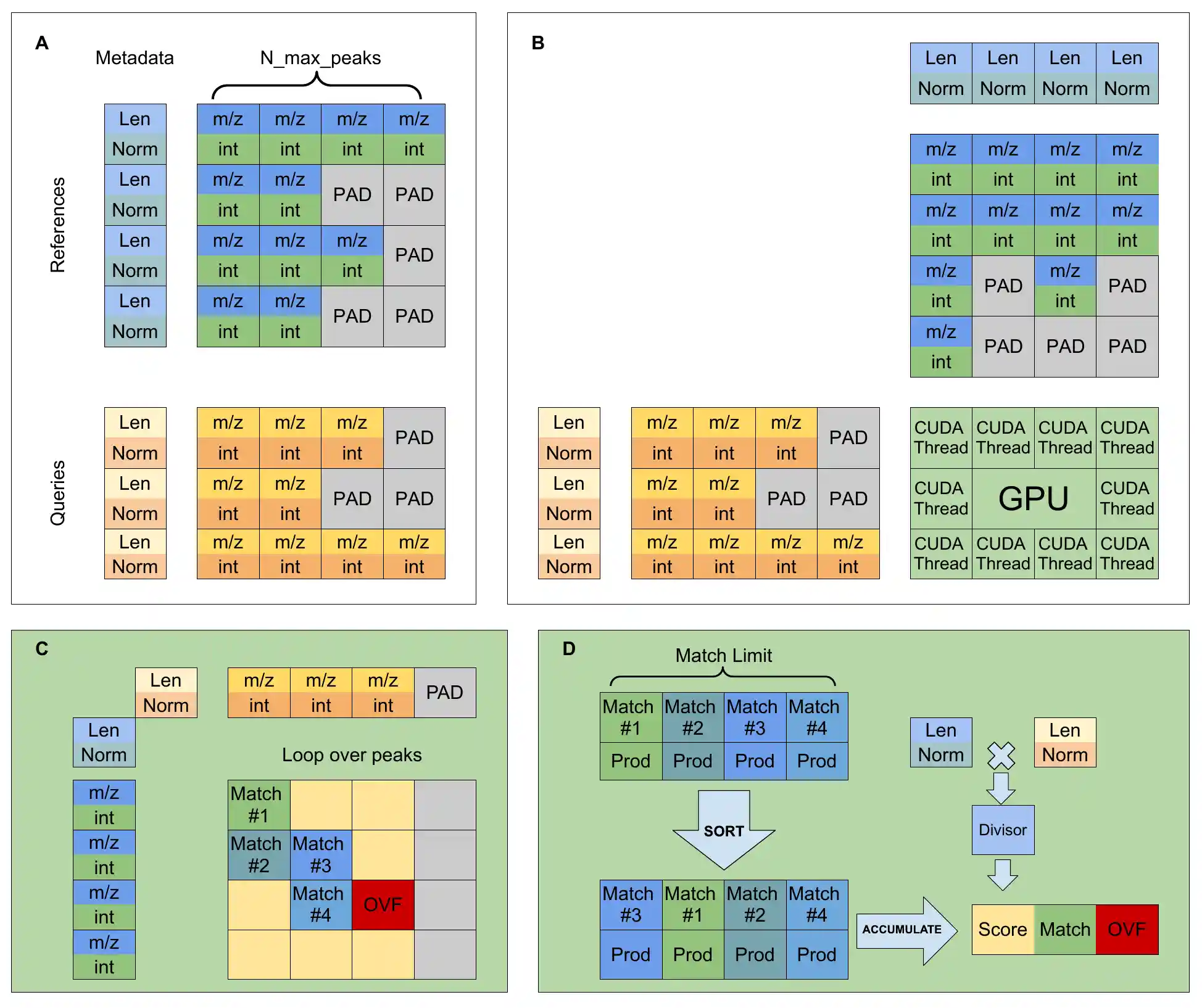
Comparing large sets of mass spectra can be done in parallel since scores can be calculated independently of each other. By leveraging a large number of threads in a GPU, we created a GPU program (kernel) that calculates a 4096x4096 similarity matrix in a fraction of a second. By iteratively calculating similarities for batches of spectra, SimMS can quickly process datasets much larger than the GPU's memory. For details, visit the preprint.
Quickstart
Hardware
Any GPU supported by Numba can be used. We tested a number of GPUs:
- GTX 1050 Ti
- RTX 2070
- T4 GPU (free on Google Colab)
- RTX 4090 (on vast.ai)
- A100 80GB (on vast.ai)
- H100SXM 80GB (on vast.ai)
Performance (comparisons/s) is proportional to the peak GPU bandwidth (GB/s).
The pytorch/pytorch:2.2.1-cuda12.1-cudnn8-devel docker image was used for development and testing.
Install
pip install git+https://github.com/PangeAI/simms
Use with MatchMS
from matchms import calculate_scores from matchms.importing import load_from_mgf from simms.utils import download from simms.similarity import CudaCosineGreedy, \ CudaModifiedCosine, \ CudaFingerprintSimilarity sample_file = download('pesticides.mgf') references = list(load_from_mgf(sample_file)) queries = list(load_from_mgf(sample_file)) similarity_function = CudaCosineGreedy() scores = calculate_scores( references=references, queries=queries, similarity_function=similarity_function, ) scores.scores_by_query(queries[42], 'CudaCosineGreedy_score', sort=True)
Supported similarity functions
-
CudaModifiedCosine, equivalent to ModifiedCosine -
CudaCosineGreedy, equivalent to CosineGreedy -
CudaFingerprintSimilarity, equivalent to FingerprintSimilarity (jaccard,cosine,dice) -
More coming soon - requests are welcome!
Installation
The easiest way to get started is to use the colab notebook that has everything ready for you.
For local installations, we recommend using micromamba, it is much faster.
Total size of install in a fresh conda environment will be around 7-8GB (heaviest packages are pytorch, and cudatoolkit).
# Install cudatoolkit conda install nvidia::cuda-toolkit -y # Install torch (follow the official guide https://pytorch.org/get-started/locally/#start-locally) conda install pytorch -c pytorch -c nvidia -y # Install numba (follow the offical guide: https://numba.pydata.org/numba-doc/latest/user/installing.html#installing-using-conda-on-x86-x86-64-power-platforms) conda install numba -y # Install this repository pip install git+https://github.com/PangeAI/simms
Run in docker
The pytorch/pytorch:2.2.1-cuda12.1-cudnn8-devel has nearly everything you need. Once inside, do:
pip install git+https://github.com/PangeAI/simms
Run on vast.ai
Use this template as a starting point, once inside, simply do:
pip install git+https://github.com/PangeAI/simms
Frequently asked questions
I want to get referenece_id, query_id and score as 1D arrays, separately. How do I do this?
Use the "sparse" mode. It directly gives you the columns. You can set sparse_threshold to 0, at which point you will get all the scores.
from simms.similarity import CudaCosineGreedy scores_cu = CudaCosineGreedy( sparse_threshold=0.75, # anything with a lower score gets discarded ).matrix(references, queries, array_type='sparse') # Unpack sparse results as 1D arrays ref_id, query_id, scores = scores_cu.data['sparse_score'] ref_id, query_id, matches = scores_cu.data['sparse_matches']
Citing SimMS
@article{onoprishvili2025simms, author = {Onoprishvili, Tornike and Yuan, Jui-Hung and Petrov, Kamen and Ingalalli, Vijay and Khederlarian, Lila and Leuchtenmuller, Niklas and Chandra, Sona and Duarte, Aurelien and Bender, Andreas and Gloaguen, Yoann}, title = {SimMS: A GPU-Accelerated Cosine Similarity implementation for Tandem Mass Spectrometry}, journal = {Bioinformatics}, year = {2025}, month = {02}, doi = {10.1093/bioinformatics/btaf081} }