General
SasView is a Small Angle Scattering (SAS) analysis package for the analysis of 1D and 2D scattering data directly in inverse space. The focus was originally on neutron data (SANS) but has been used for X-rays as well and includes a tool for determining a slit resolution for the SAXSess instrument. SasView also includes PrView to invert SAS data to P(r), a resolution calculator, and a scattering length density calculator among others tools. A simple plugin mechanism is available for users to add custom models.
Acknowledgement
This project was initiated by the NSF-funded DANSE project, DMR-0520547, a SANS sub-project at the University of Tennessee. Acknowledgement of that original funding would be appreciated in any publications that make use of the software.
This project received funding from the European Union’s Horizon 2020 research and innovation programme under the SINE2020 project, grant agreement No 654000.
The latest stable releases of SasView can be found on the website. The following instructions will focus on providing materials for scripting, developing and would lead you to the most important resources.
Install instructions
Users
Users can install SasView either from the installers listed on the SasView website,
or directly from the packages distributed by pip.
Install using pip inside a virtual environment in the current directory:
python -m venv .venv # create the environment . .venv/bin/activate # activate the environment on linux and MacOS # .venv\Scripts\activate & REM Windows: activate environment python -m pip install sasview python -m sas # launch the gui
Install using uv for your user:
uv tool install sasview
uvx sasview # launch the guiNote: To launch SasView, it needs to be installed. Running SasView from a source directory is not supported.
Developers
The installation instructions for developers can be found here.
Installing the development-version of SasView with conda is currently not supported.
NOTE: In case you want to contribute, please also checkout the DevelopersNotes.
Getting Started
Scripting
This section is a small scripting example in SasView to check your installation. We will fit a simple sphere model. For this first lets synthesize input data.
import numpy as np from sasmodels.bumps_model import Model from sasmodels.core import load_model from sasmodels.direct_model import call_kernel # define q vector q = np.logspace(-3, -0.1, 200) # define the model exp_model = load_model("sphere") exp_pars = { "radius": 50, "sld": 1, "sld_solvent": 6, "scale": 1, "background": 0.001, } # calculate intensities Iq = call_kernel(exp_model.make_kernel([q]), exp_pars) # calculate errors and normalize data max_counts = 1e7 # approximate number of counts at first q values norm = Iq[0] / max_counts counts = np.random.poisson((Iq / norm).astype(int)) errors = np.sqrt(counts) * norm data = counts * norm dataset = np.array([q, data, errors]).T # saving the data header = ( "Neutron-like data generated for model " + exp_model.info.name + " with parameters:\n" ) for key in exp_pars.keys(): header += key + " = " + str(exp_pars[key]) + "\n" header += "Q\t counts\t error" np.savetxt("scattering.txt", dataset, fmt="%12.6e", delimiter="\t", header=header)
...and now let's do the fitting. We will optimize the scale, radius and background starting from an inital values close to the ground truth.
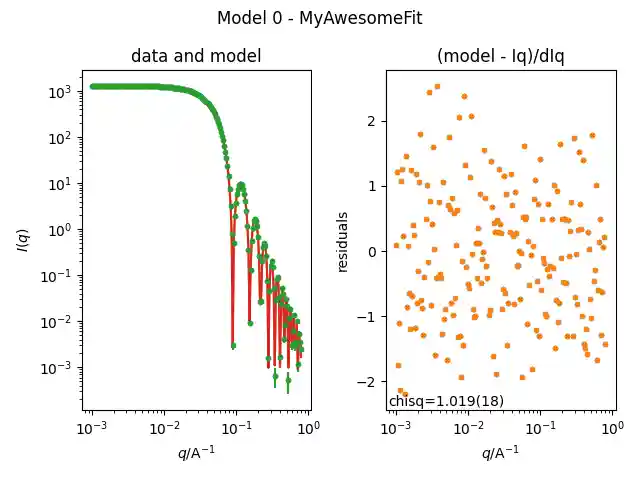
import matplotlib.pyplot as plt from sasmodels.bumps_model import Model, Experiment from sasmodels.core import load_model from sasmodels.data import load_data from bumps.fitters import fit from bumps.names import FitProblem # defining the model to fit fit_pars = { "radius": 80, "sld": 1, "sld_solvent": 6, "scale": 0.900, "background": 0.05, } fit_kernel = load_model("sphere") fit_model = Model(fit_kernel, **fit_pars) ## setting fitting ranges fit_model.radius.range(10, 1000) fit_model.scale.range(1e-3, 10) fit_model.background.range(1e-9, 0.1) # load the data we synthesized above exp_data = load_data("scattering.txt") # Setup the experiments, sharing the same model across all datasets. M = Experiment(data=exp_data, model=fit_model, name="MyAwsomeFit") problem = FitProblem(M) plt.figure() problem.plot(view=True) # fit the results result = fit(problem, method="dream") print(f"Final chisq {problem.chisq()}\n") problem.plot() for k, v, dv in zip(problem.labels(), result.x, result.dx): print(f"{k} : {v:.4f} +- {dv:.4f}") plt.show()
This simple fit should results in a $\chi^2$ close to one.
Resources
In case you are just getting started or you want to contribute please checkout some selected resources.