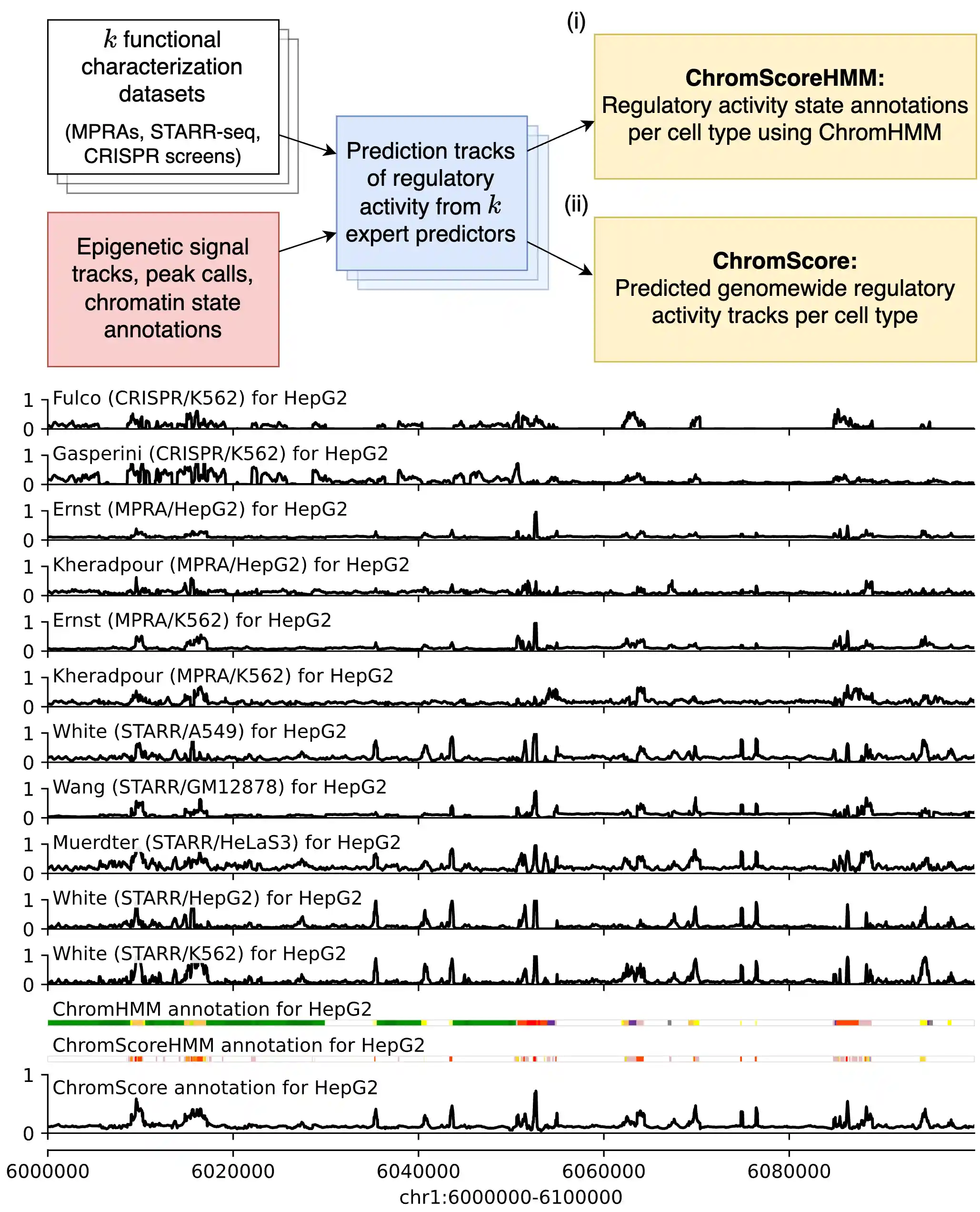
ChromActivity is a computational framework for the annotation of regulatory activity genomewide, through integration of data from epigenomic maps and multiple functional characterization assays.
We generate three types of annotations:
- Expert score tracks: Genomewide regulatory activity prediction tracks associated with each functional characterization assay dataset across all cell and tissue types with epigenome data
- ChromScoreHMM annotations: Cell type specific genome annotations based on the combinatorial and spatial patterns within the expert predictions
- ChromScore tracks: Genomewide, cell type specific ensemble regulatory activity prediction tracks. Provides a numerical score for each 25 bp interval in the genome
Precomputed annotations
Precomputed annotations are available to download at: https://ucla.box.com/v/chromactivity
View annotations on the UCSC Genome Browser: session link, track hub link
Getting started
ChromActivity manages its dependencies using the conda package manager. Mambaforge is the recommended distribution for installing conda.
# Download ChromActivity from repository git clone --depth 1 https://github.com/ernstlab/chromactivity # Set up conda environment cd chromactivity conda env create -f environment.yml conda activate chromactivity_env # Download and extract ChromHMM wget -N -P vendored https://ernstlab.biolchem.ucla.edu/ChromHMM/ChromHMM.zip unzip -o vendored/ChromHMM.zip -d vendored
Raw data directories
By default, ChromActivity uses imputed epigenomic data from the Roadmap Epigenomics compendium with the following directory structure:
# Data URLs: https://egg2.wustl.edu/roadmap/web_portal/imputed.html # Imputed signal tracks f"data/raw/roadmap/signal/{cell_type}/{cell_type}-{mark}.imputed.pval.signal.bigwig" # Peak calls f"data/raw/roadmap/peaks/{cell_type}/{cell_type}-{mark}.imputed.narrowPeak.bed.nPk.gz" # 25-State ChromHMM model f"data/raw/roadmap/chromstate/chromstate_25/{cell_type}/{cell_type}_25_imputed12marks_mnemonics.sorted.bed",
Overriding the default directory structure is possible by modifying chromactivity/mappings.py.
Usage examples
Command line usage examples:
# Train and serialize ChromActivity experts using the default labels in "data/labels" chromactivity train_experts --labels_dir data/labels --model_out_fn "models/my_chromactivity.model" # Generate tracks from serialized model for the HepG2 (Roadmap epigenome ID: E118) cell type chromactivity generate_tracks --model_fn "models/chromactivity.model" --cell_types "E118" --coords_bed_fn "data/external/test.bed" --combined_bigwigs_out_dir "tracks/" # Generate ChromScoreHMM annotations from generated tracks chromactivity train_chromscorehmm --num_states 15 --track_dir "tracks/" --out-dir "models/chromscorehmm"
External resources
vendored/ChromHMM/ChromHMM.jar: https://ernstlab.biolchem.ucla.edu/ChromHMMdata/external/hg19.chrom.sizes: https://hgdownload.soe.ucsc.edu/goldenPath/hg19/bigZips/hg19.chrom.sizes
References
Dincer, T., Ernst, J. ChromActivity: integrative epigenomic and functional characterization assay based annotation of regulatory activity across diverse human cell types. Genome Biol 26, 123 (2025). https://doi.org/10.1186/s13059-025-03579-6