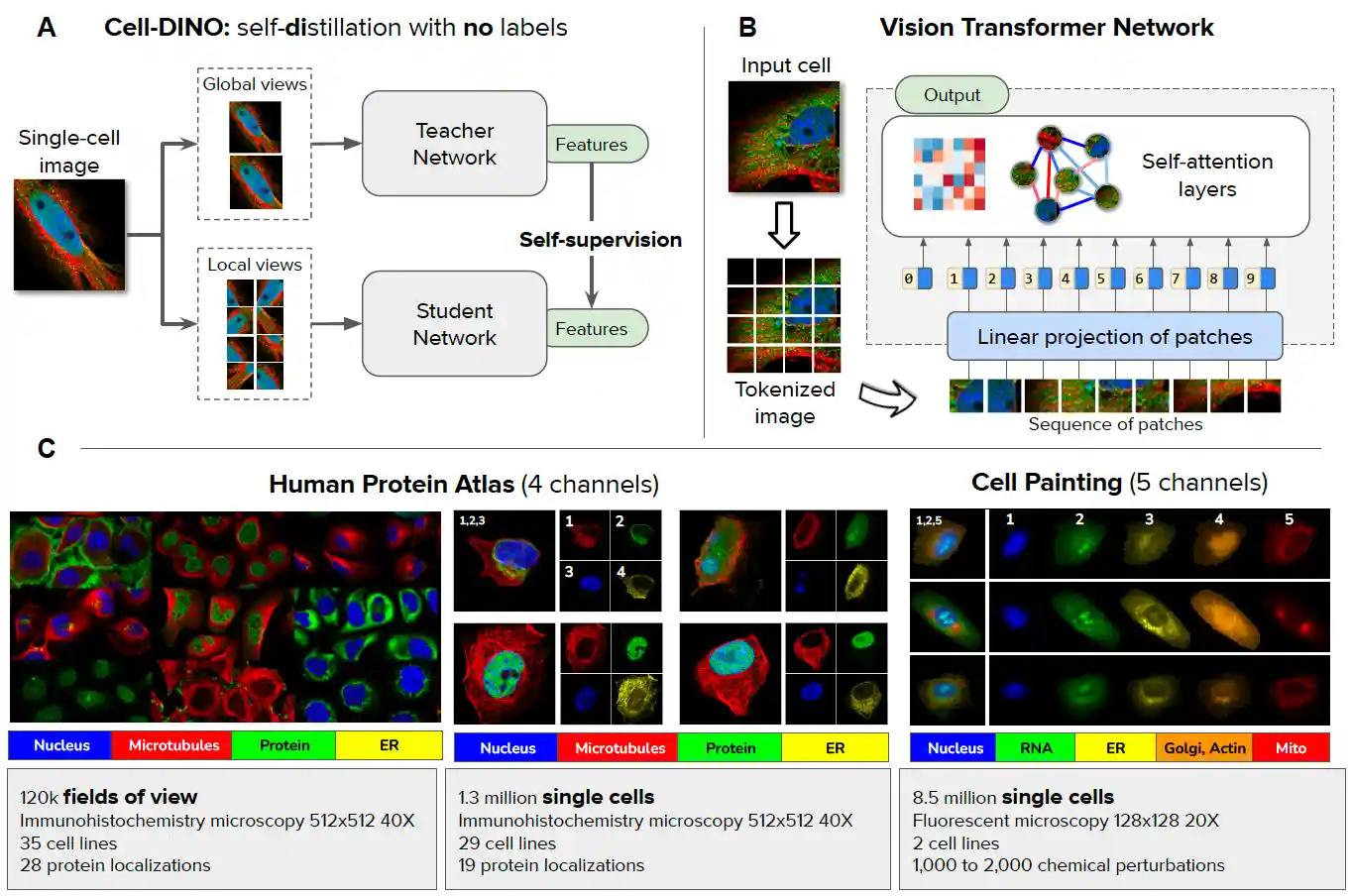
Cell-DINO: Self-Supervised Image-based Embeddings for Cell Fluorescent Microscopy
Théo Moutakanni*, Camille Couprie*, Seungeun Yi*, Elouan Gardes*, Piotr Bojanowski*, Hugo Touvron*, Michael Doron, Zitong S. Chen, Nikita Moshkov, Mathilde Caron, Armand Joulin, Wolfgang M. Pernice, Juan C. Caicedo
[BibTeX]
PyTorch implementation and pretrained models for Cell-DINO.
The contents of this repo, including the code and model weights, are intended for research use only. It is not for use in medical procedures, including any diagnostics, treatment, or curative applications. Do not use this model for any clinical purpose or as a substitute for professional medical judgement.
Pretrained models
One model pretrained on HPA single cell, one model pretrained on HPA Field of View, one model pretrained on HPA Field of View at a larger resolution and one model pretrained on the combined cell painting dataset are released, see dowload instructions at the end of README.md, section "DINO for Biology".
Installation
Follow instructions in the DINOv2 README or build the following environment:
conda create -n py310 python=3.10 conda activate py310 pip install -r requirements.txt pip install -U scikit-learn
Data preparation
The HPA-FoV and HPA single cell (HPAone) datasets are available here The folder also contains three sample images ("sample_images") that can be dowloaded to test quickly inference in the provided notebook in notebooks/cell_dino
THe required files for HPA single cell (dataloader : HPAone.py) are:
fixed_size_masked_single_cells_HPA varied_size_masked_single_cells_HPA varied_size_masked_single_cells_pretrain_20240507.csv fixed_size_masked_single_cells_evaluation_20240507.csv fixed_size_masked_single_cells_pretrain_20240507.csv
⚠️ To execute the commands provided in the next sections for training and evaluation, the dinov2 package should be included in the Python module search path, i.e. simply prefix the command to run with PYTHONPATH=..
Training
Fast setup: training DINOv2 ViT-L/16 on HPA single cell dataset
Run CellDINO training on 4 A100-80GB nodes (32 GPUs) in a SLURM cluster environment with submitit:
python dinov2/run/train/train.py \
--nodes 4 \
--config-file dinov2/configs/train/cell_dino/vitl16_hpaone.yaml \
--output-dir <PATH/TO/OUTPUT/DIR> \
train.dataset_path=HPAone:split=ALL:root=<PATH/TO/DATASET>Training time is approximately 2 days on 4 A100 GPU nodes and the resulting checkpoint should reach 78.5 F1 accuracy for protein localization with a linear evaluation.
The training code saves the weights of the teacher in the eval folder every 9000 iterations for evaluation.
Evaluation
The training code regularly saves the teacher weights. In order to evaluate the model, run the following evaluation on a single node:
Linear classification with data augmentation on HPAone or HPAFov (replace 'HPAone' by 'HPAFoV'):
PYTHONPATH=.:dinov2/data python dinov2/run/eval/cell_dino/linear.py \ --config-file dinov2/configs/eval/cell_dino/vitl16_pretrain.yaml \ --pretrained-weights <PATH/TO/OUTPUT/DIR>/eval/training_44999/teacher_checkpoint.pth \ --output-dir <PATH/TO/OUTPUT/DIR>/eval/training_44999/linear \ --train-dataset HPAone:split=TRAIN:mode=PROTEIN_LOCALIZATION:root=<PATH/TO/DATASET> \ --val-dataset HPAone:split=VAL:mode=PROTEIN_LOCALIZATION:root<PATH/TO/DATASET> \ --val-metric-type mean_per_class_multilabel_f1 \ --loss-type binary_cross_entropy \ --avgpool \
We release the weights from evaluating the different models:
The performance of the provided pretrained model weights can be evaluated as follows on HPAone and HPAFoV (replace 'HPAone' by 'HPAFoV', and make sure to load the correct model weights) for the protein localization task:
PYTHONPATH=.:dinov2/data python dinov2/run/eval/linear_celldino.py \ --config-file dinov2/configs/eval/cell_dino/vitl16_pretrain.yaml \ --pretrained-weights <CHECKPOINT/PATH> \ --output-dir <PATH/TO/OUTPUT/DIR> \ --train-dataset HPAone:split=TRAIN:mode=PROTEIN_LOCALIZATION:root=<PATH/TO/DATASET> \ --val-dataset HPAone:split=VAL:mode=PROTEIN_LOCALIZATION:root=root<PATH/TO/DATASET> \ --val-metric-type mean_per_class_multilabel_f1 \ --loss-type binary_cross_entropy \ --avgpool \
and
PYTHONPATH=.:dinov2/data python dinov2/run/eval/cell_dino/linear.py \ --config-file dinov2/configs/eval/cell_dino/vitl16_pretrain.yaml \ --pretrained-weights <CHECKPOINT/PATH> \ --output-dir <PATH/TO/OUTPUT/DIR> \ --train-dataset HPAone:split=TRAIN:mode=CELL_TYPE:root=<PATH/TO/DATASET> \ --val-dataset HPAone:split=VAL:mode=CELL_TYPE:root=root<PATH/TO/DATASET> \ --val-metric-type mean_per_class_multiclass_f1 \ --avgpool \
for the cell line classification task.
knn evaluation on HPAone:
PYTHONPATH=.:dinov2/data python dinov2/run/eval/cell_dino/knn.py \ --config-file dinov2/configs/eval/cell_dino/vitl16_pretrain.yaml \ --pretrained-weights <CHECKPOINT/PATH>\ --output-dir <PATH/TO/OUTPUT/DIR>\ --train-dataset HPAone:split=TRAIN:mode=CELL_TYPE:root=<PATH/TO/DATASET> \ --val-dataset HPAone:split=VAL:mode=CELL_TYPE:root=<PATH/TO/DATASET> \ --metric-type mean_per_class_multiclass_f1 \ --crop-size 384 \ --batch-size 256 \ --resize-size 0 \ --nb_knn 1 \
for the cell line classification task. For the knn evaluation on HPAFoV, replace 'HPAone' by 'HPAFoV' in the command above. For protein localization, don't forget to change the metric type to mean_per_class_multilabel_f1.
License
Cell-DINO code is released under the CC by NC licence See LICENSE_CELL_DINO_CODE for additional details. Model weights will be released under the FAIR Non-Commercial Research License. See LICENSE_CELL_DINO_MODELS for additional details.
Contributing
See contributing and the code of conduct.
Citing Cell-DINO
If you find this repository useful, please consider giving a star ⭐ and citation 🦖:
@misc{,
title={Cell-DINO: Self-Supervised Image-based Embeddings for Cell Fluorescent Microscopy},
author={Moutakanni, Th\'eo and Couprie, Camille and Yi, Seungeun and Gardes, Elouan Gardes and Bojanowski, Piotr and Touvron, Hugo and Doron, Michael and Chen, Zitong S. and Moshkov, Nikita and Caron, Mathilde and Joulin, Armand and Pernice, Wolfgang M. and Caicedo, Juan C.},
journal={in review to PloS One on Computational Biology},
year={2025}
}