Implementation of the Robust Random Cut Forest Algorithm for anomaly detection by Guha et al. (2016).
S. Guha, N. Mishra, G. Roy, & O. Schrijvers, Robust random cut forest based anomaly detection on streams, in Proceedings of the 33rd International conference on machine learning, New York, NY, 2016 (pp. 2712-2721).
About
The Robust Random Cut Forest (RRCF) algorithm is an ensemble method for detecting outliers in streaming data. RRCF offers a number of features that many competing anomaly detection algorithms lack. Specifically, RRCF:
- Is designed to handle streaming data.
- Performs well on high-dimensional data.
- Reduces the influence of irrelevant dimensions.
- Gracefully handles duplicates and near-duplicates that could otherwise mask the presence of outliers.
- Features an anomaly-scoring algorithm with a clear underlying statistical meaning.
This repository provides an open-source implementation of the RRCF algorithm and its core data structures for the purposes of facilitating experimentation and enabling future extensions of the RRCF algorithm.
Documentation
Read the docs here 📖.
Installation
Use pip to install rrcf via pypi:
Currently, only Python 3 is supported.
Dependencies
The following dependencies are required to install and use rrcf:
- numpy (>= 1.15)
The following optional dependencies are required to run the examples shown in the documentation:
- pandas (>= 0.23)
- scipy (>= 1.2)
- scikit-learn (>= 0.20)
- matplotlib (>= 3.0)
Listed version numbers have been tested and are known to work (this does not necessarily preclude older versions).
Robust random cut trees
A robust random cut tree (RRCT) is a binary search tree that can be used to detect outliers in a point set. A RRCT can be instantiated from a point set. Points can also be added and removed from an RRCT.
Creating the tree
import numpy as np import rrcf # A (robust) random cut tree can be instantiated from a point set (n x d) X = np.random.randn(100, 2) tree = rrcf.RCTree(X) # A random cut tree can also be instantiated with no points tree = rrcf.RCTree()
Inserting points
tree = rrcf.RCTree() for i in range(6): x = np.random.randn(2) tree.insert_point(x, index=i)
─+
├───+
│ ├───+
│ │ ├──(0)
│ │ └───+
│ │ ├──(5)
│ │ └──(4)
│ └───+
│ ├──(2)
│ └──(3)
└──(1)
Deleting points
─+
├───+
│ ├───+
│ │ ├──(0)
│ │ └───+
│ │ ├──(5)
│ │ └──(4)
│ └──(3)
└──(1)
Anomaly score
The likelihood that a point is an outlier is measured by its collusive displacement (CoDisp): if including a new point significantly changes the model complexity (i.e. bit depth), then that point is more likely to be an outlier.
# Seed tree with zero-mean, normally distributed data X = np.random.randn(100,2) tree = rrcf.RCTree(X) # Generate an inlier and outlier point inlier = np.array([0, 0]) outlier = np.array([4, 4]) # Insert into tree tree.insert_point(inlier, index='inlier') tree.insert_point(outlier, index='outlier')
tree.codisp('inlier') >>> 1.75
tree.codisp('outlier') >>> 39.0
Batch anomaly detection
This example shows how a robust random cut forest can be used to detect outliers in a batch setting. Outliers correspond to large CoDisp.
import numpy as np import pandas as pd import rrcf # Set parameters np.random.seed(0) n = 2010 d = 3 num_trees = 100 tree_size = 256 # Generate data X = np.zeros((n, d)) X[:1000,0] = 5 X[1000:2000,0] = -5 X += 0.01*np.random.randn(*X.shape) # Construct forest forest = [] while len(forest) < num_trees: # Select random subsets of points uniformly from point set ixs = np.random.choice(n, size=(n // tree_size, tree_size), replace=False) # Add sampled trees to forest trees = [rrcf.RCTree(X[ix], index_labels=ix) for ix in ixs] forest.extend(trees) # Compute average CoDisp avg_codisp = pd.Series(0.0, index=np.arange(n)) index = np.zeros(n) for tree in forest: codisp = pd.Series({leaf : tree.codisp(leaf) for leaf in tree.leaves}) avg_codisp[codisp.index] += codisp np.add.at(index, codisp.index.values, 1) avg_codisp /= index
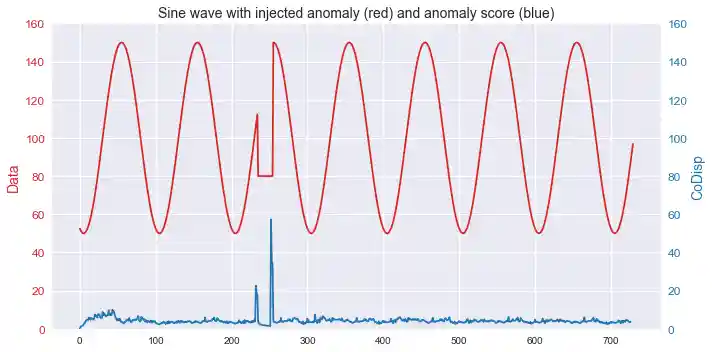
Streaming anomaly detection
This example shows how the algorithm can be used to detect anomalies in streaming time series data.
import numpy as np import rrcf # Generate data n = 730 A = 50 center = 100 phi = 30 T = 2*np.pi/100 t = np.arange(n) sin = A*np.sin(T*t-phi*T) + center sin[235:255] = 80 # Set tree parameters num_trees = 40 shingle_size = 4 tree_size = 256 # Create a forest of empty trees forest = [] for _ in range(num_trees): tree = rrcf.RCTree() forest.append(tree) # Use the "shingle" generator to create rolling window points = rrcf.shingle(sin, size=shingle_size) # Create a dict to store anomaly score of each point avg_codisp = {} # For each shingle... for index, point in enumerate(points): # For each tree in the forest... for tree in forest: # If tree is above permitted size, drop the oldest point (FIFO) if len(tree.leaves) > tree_size: tree.forget_point(index - tree_size) # Insert the new point into the tree tree.insert_point(point, index=index) # Compute codisp on the new point and take the average among all trees if not index in avg_codisp: avg_codisp[index] = 0 avg_codisp[index] += tree.codisp(index) / num_trees
Obtain feature importance
This example shows how to estimate the feature importance using the dimension of cut obtained during the calculation of the CoDisp.
import numpy as np import pandas as pd import rrcf # Set parameters np.random.seed(0) n = 2010 d = 3 num_trees = 100 tree_size = 256 # Generate data X = np.zeros((n, d)) X[:1000,0] = 5 X[1000:2000,0] = -5 X += 0.01*np.random.randn(*X.shape) # Construct forest forest = [] while len(forest) < num_trees: # Select random subsets of points uniformly from point set ixs = np.random.choice(n, size=(n // tree_size, tree_size), replace=False) # Add sampled trees to forest trees = [rrcf.RCTree(X[ix], index_labels=ix) for ix in ixs] forest.extend(trees) # Compute average CoDisp with the cut dimension for each point dim_codisp = np.zeros([n,d],dtype=float) index = np.zeros(n) for tree in forest: for leaf in tree.leaves: codisp,cutdim = tree.codisp_with_cut_dimension(leaf) dim_codisp[leaf,cutdim] += codisp index[leaf] += 1 avg_codisp = dim_codisp.sum(axis=1)/index #codisp anomaly threshold and calculate the mean over each feature feature_importance_anomaly = np.mean(dim_codisp[avg_codisp>50,:],axis=0) #create a dataframe with the feature importance df_feature_importance = pd.DataFrame(feature_importance_anomaly,columns=['feature_importance']) df_feature_importance
Contributing
We welcome contributions to the rrcf repo. To contribute, submit a pull request to the dev branch.
Types of contributions
Some suggested types of contributions include:
- Bug fixes
- Documentation improvements
- Performance enhancements
- Extensions to the algorithm
Check the issue tracker for any specific issues that need help. If you encounter a problem using rrcf, or have an idea for an extension, feel free to raise an issue.
Guidelines for contributors
Please consider the following guidelines when contributing to the codebase:
- Ensure that any new methods, functions or classes include docstrings. Docstrings should include a description of the code, as well as descriptions of the inputs (arguments) and outputs (returns). Providing an example use case is recommended (see existing methods for examples).
- Write unit tests for any new code and ensure that all tests are passing with no warnings. Please ensure that overall code coverage does not drop below 80%.
Running unit tests
To run unit tests, first ensure that pytest and pytest-cov are installed:
$ pip install pytest pytest-cov
To run the tests, navigate to the root directory of the repo and run:
Citing
If you have used this codebase in a publication and wish to cite it, please use the Journal of Open Source Software article.
M. Bartos, A. Mullapudi, & S. Troutman, rrcf: Implementation of the Robust Random Cut Forest algorithm for anomaly detection on streams, in: Journal of Open Source Software, The Open Journal, Volume 4, Number 35. 2019
@article{bartos_2019_rrcf, title={{rrcf: Implementation of the Robust Random Cut Forest algorithm for anomaly detection on streams}}, authors={Matthew Bartos and Abhiram Mullapudi and Sara Troutman}, journal={{The Journal of Open Source Software}}, volume={4}, number={35}, pages={1336}, year={2019} }