Foundation Model for Endoscopy Video Analysis
🔥 You may also check our improved version of this foundation model for handling much longer endoscopy video sequences at EndoFM-LV.
This repository provides the official PyTorch implementation of the paper Foundation Model for Endoscopy Video Analysis via Large-scale Self-supervised Pre-train by Zhao Wang*, Chang Liu*, Shaoting Zhang†, and Qi Dou†.
Key Features
- First foundation model for endoscopy video analysis.
- A large-scale endoscopic video dataset with over 33K video clips.
- Support 3 types of downstream tasks, including classification, segmentation, and detection.
Links
Details
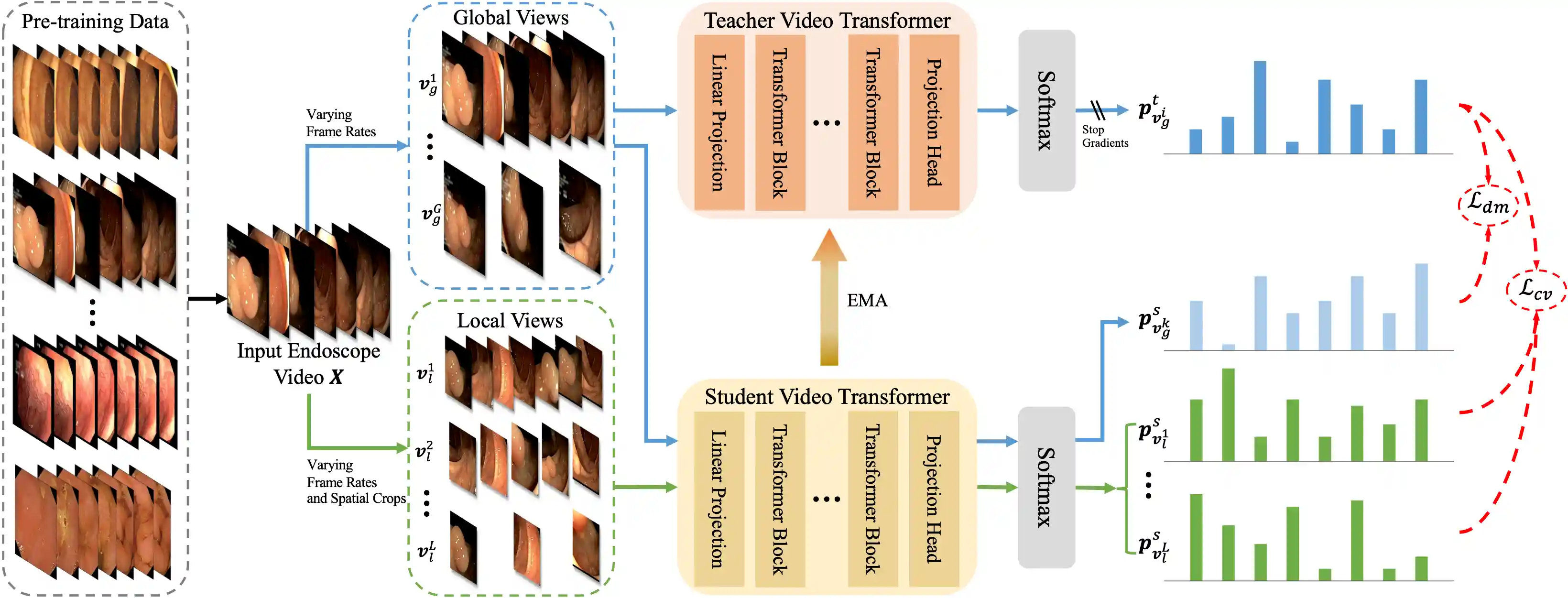
Foundation models have exhibited remarkable success in various applications, such as disease diagnosis and text report generation. To date, a foundation model for endoscopic video analysis is still lacking. In this paper, we propose Endo-FM, a foundation model specifically developed using massive endoscopic video data. First, we build a video transformer, which captures both local and global long-range dependencies across spatial and temporal dimensions. Second, we pre-train our transformer model using global and local views via a self-supervised manner, aiming to make it robust to spatial-temporal variations and discriminative across different scenes. To develop the foundation model, we construct a large-scale endoscopy video dataset by combining 9 publicly available datasets and a privately collected dataset from Baoshan Branch of Renji Hospital in Shanghai, China. Our dataset overall consists of over 33K video clips with up to 5 million frames, encompassing various protocols, target organs, and disease types. Our pre-trained Endo-FM can be easily adopted for a given downtream task via fine-tuning by serving as the backbone. With experiments on 3 different types of downstream tasks, including classification, segmentation, and detection, our Endo-FM surpasses the current state-of-the-art self-supervised pre-training and adapter-based transfer learning methods by a significant margin.
Datasets
We utilize 6 public and 1 private datasets for pre-training and 3 datasets as the downstream tasks. Except for SUN & SUN-SEG, we provide our preprocessed data for pre-training and downstream tasks.
Pre-training Data (6 public + 1 private)
- Colonoscopic [original paper] [original dataset] [our preprocessed dataset]
- SUN & SUN-SEG [original paper1] [original paper2] [original dataset1] [original dataset2]
- LPPolypVideo [original paper] [original dataset] [our preprocessed dataset]
- Hyper-Kvasir [original paper] [original dataset] [our preprocessed dataset]
- Kvasir-Capsule [original paper] [original dataset] [our preprocessed dataset]
- CholecTriplet [original paper] [original dataset] [our preprocessed dataset]
- Our Private [our preprocessed dataset]
Downstream Data (3 public)
- PolypDiag [original paper] [original dataset] [our preprocessed dataset]
- CVC-12k [original paper] [original dataset] [our preprocessed dataset]
- KUMC [original paper] [original dataset] [our preprocessed dataset]
For SUN & SUN-SEG, you need first request the original videos following this instruction. Then, you can transfer the data for pre-training videos by the following:
cd Endo-FM/data
python sun.py
python sun_seg.py
python trans_videos_pretrain.pyFinally, generating the video list pretrain/train.csv for pre-training by the following:
cd Endo-FM/data
python gencsv.pyGet Started
Main Requirements
- torch==1.8.0
- torchvision==0.9.0
- pillow==6.2.2
- timm==0.4.12
Installation
We suggest using Anaconda to setup environment on Linux, if you have installed anaconda, you can skip this step.
wget https://repo.anaconda.com/archive/Anaconda3-2020.11-Linux-x86_64.sh && zsh Anaconda3-2020.11-Linux-x86_64.shThen, we can install packages using provided environment.yaml.
cd Endo-FM
conda env create -f environment.yaml
conda activate endofmPre-trained Weights
You can directly download our pre-trained Endo-FM via this link and put it under checkpoints/ for downstream fine-tuning.
Downstream Fine-tuned Weights
Also, we provide the pre-trained weights of 3 downstream tasks for direct downstream testing.
| Dataset | PolypDiag | CVC-12k | KUMC |
|---|---|---|---|
| Our Paper | 90.7 | 73.9 | 84.1 |
| Released Model | 91.5 | 76.6 | 84.0 |
| Weights | link | link | link |
Pre-training
cd Endo-FM
wget -P checkpoints/ https://github.com/kahnchana/svt/releases/download/v1.0/kinetics400_vitb_ssl.pth
bash scripts/train_clips32k.shDownstream Fine-tuning
# PolypDiag (Classification) cd Endo-FM bash scripts/eval_finetune_polypdiag.sh # CVC (Segmentation) cd Endo-FM/TransUNet python train.py # KUMC (Detection) cd Endo-FM/STMT python setup.py build develop python -m torch.distributed.launch \ --nproc_per_node=1 \ tools/train_net.py \ --master_port=$((RANDOM + 10000)) \ --config-file configs/STFT/kumc_R_50_STFT.yaml \ OUTPUT_DIR log_dir/kumc_finetune
Direct Downstream Testing
# PolypDiag (Classification) cd Endo-FM bash scripts/test_finetune_polypdiag.sh # CVC (Segmentation) cd Endo-FM/TransUNet python train.py --test # KUMC (Detection) cd Endo-FM/STMT python setup.py build develop python -m torch.distributed.launch \ --nproc_per_node=1 \ tools/test_net.py \ --master_port=$((RANDOM + 10000)) \ --config-file configs/STFT/kumc_R_50_STFT.yaml \ MODEL.WEIGHT kumc.pth \ OUTPUT_DIR log_dir/kumc_finetune
🙋♀️ Feedback and Contact
For further questions, pls feel free to contact Zhao Wang.
🛡️ License
This project is under the Apache License 2.0 license. See LICENSE for details.
🙏 Acknowledgement
Our code is based on DINO, TimeSformer, SVT, TransUNet, and STFT. Thanks them for releasing their codes.
📝 Citation
If you find this code useful, please cite in your research papers.
@inproceedings{
wang2023foundation,
title={Foundation Model for Endoscopy Video Analysis via Large-scale Self-supervised Pre-train},
author={Zhao Wang and Chang Liu and Shaoting Zhang and Qi Dou},
booktitle={International Conference on Medical Image Computing and Computer-Assisted Intervention},
pages={101--111},
year={2023},
organization={Springer}
}