Python package containing tools for radial basis function (RBF) applications. Applications include interpolating/smoothing scattered data and solving PDEs over irregular domains. The complete documentation for this package can be found here.
Features
- Functions for evaluating RBFs and their exact derivatives.
- A class for RBF interpolation, which is used for interpolating and smoothing scattered, noisy, N-dimensional data.
- An abstraction for Gaussian processes. Gaussian processes are primarily used here for Gaussian process regression (GPR), which is a nonparametric Bayesian interpolation/smoothing method.
- An algorithm for generating Radial Basis Function Finite Difference (RBF-FD) weights. This is used for solving large scale PDEs over irregular domains.
- A node generation algorithm which can be used for solving PDEs with the spectral RBF method or the RBF-FD method.
- Halton sequence generator.
- Computational geometry functions (e.g. point in polygon testing) for 1, 2, and 3 spatial dimensions.
Installation
RBF requires the following python packages: numpy, scipy, sympy, cython, and networkx. These dependencies should be satisfied with just the base Anaconda python package (https://www.continuum.io/downloads)
download the RBF package
$ git clone http://github.com/treverhines/RBF.git
compile and install
$ cd RBF
$ python setup.py installtest that everything works
$ cd test $ python -m unittest discover
Demonstration
Smoothing Scattered Data
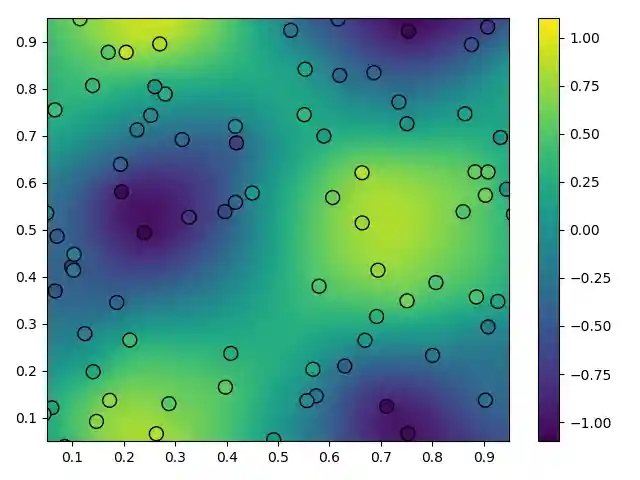
''' In this example we generate synthetic scattered data with added noise and then fit it with a smoothed RBF interpolant. The interpolant in this example is equivalent to a thin plate spline. ''' import numpy as np from rbf.interpolate import RBFInterpolant import rbf.basis import matplotlib.pyplot as plt np.random.seed(1) basis = rbf.basis.phs2 order = 1 x_obs = np.random.random((100,2)) # observation points u_obs = np.sin(2*np.pi*x_obs[:,0])*np.cos(2*np.pi*x_obs[:,1]) # signal u_obs += np.random.normal(0.0,0.2,100) # add noise to signal I = RBFInterpolant(x_obs,u_obs,penalty=0.1,basis=basis,order=order) vals = np.linspace(0,1,200) x_itp = np.reshape(np.meshgrid(vals,vals),(2,200*200)).T # interp points u_itp = I(x_itp) # evaluate the interpolant # plot the results plt.tripcolor(x_itp[:,0],x_itp[:,1],u_itp,vmin=-1.1,vmax=1.1,cmap='viridis') plt.scatter(x_obs[:,0],x_obs[:,1],s=100,c=u_obs,vmin=-1.1,vmax=1.1, cmap='viridis',edgecolor='k') plt.xlim((0.05,0.95)) plt.ylim((0.05,0.95)) plt.colorbar() plt.tight_layout() plt.savefig('../figures/interpolate.a.png') plt.show()
Plot generated by the above code. Observations are shown as scatter points and the smoothed interpolant is the color field.
Solving PDEs
There are two methods for solving PDEs with RBFs: the spectral method and the RBF-FD method. The spectral method has been touted as having remarkable accuracy; however it is only applicable for small scale problems and requires a good choice for a shape parameter. The RBF-FD method is appealing because it can be used for large scale problems, there is no need to tune a shape parameter (assuming you use polyharmonic splines to generate the weights), and higher order accuracy can be attained by simply increasing the stencil size or increasing the order of the polynomial used to generate the weights. In short, the RBF-FD method should always be preferred over the spectral RBF method. An example of the two methods is provided below.
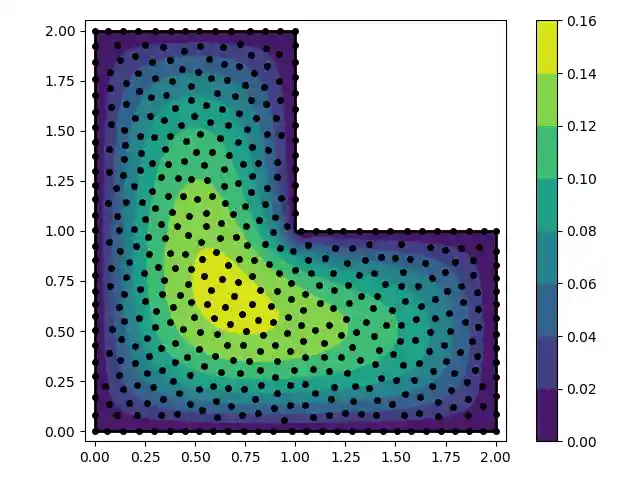
''' In this example we solve the Poisson equation over an L-shaped domain with fixed boundary conditions. We use the multiquadratic RBF (*mq*) with a shape parameter that scales inversely with the average nearest neighbor distance. ''' import numpy as np from rbf.basis import mq from rbf.geometry import contains from rbf.nodes import menodes,neighbors import matplotlib.pyplot as plt # Define the problem domain with line segments. vert = np.array([[0.0,0.0],[2.0,0.0],[2.0,1.0], [1.0,1.0],[1.0,2.0],[0.0,2.0]]) smp = np.array([[0,1],[1,2],[2,3],[3,4],[4,5],[5,0]]) N = 500 # total number of nodes nodes,smpid = menodes(N,vert,smp) # generate nodes edge_idx, = (smpid>=0).nonzero() # identify edge nodes interior_idx, = (smpid==-1).nonzero() # identify interior nodes dx = np.mean(neighbors(nodes,2)[1][:,1]) # avg. distance to nearest neighbor eps = 0.5/dx # shape parameter # create "left hand side" matrix A = np.empty((N,N)) A[interior_idx] = mq(nodes[interior_idx],nodes,eps=eps,diff=[2,0]) A[interior_idx] += mq(nodes[interior_idx],nodes,eps=eps,diff=[0,2]) A[edge_idx] = mq(nodes[edge_idx],nodes,eps=eps) # create "right hand side" vector d = np.empty(N) d[interior_idx] = -1.0 # forcing term d[edge_idx] = 0.0 # boundary condition # Solve for the RBF coefficients coeff = np.linalg.solve(A,d) # interpolate the solution on a grid xg,yg = np.meshgrid(np.linspace(-0.05,2.05,400),np.linspace(-0.05,2.05,400)) points = np.array([xg.flatten(),yg.flatten()]).T u = mq(points,nodes,eps=eps).dot(coeff) # evaluate at the interp points u[~contains(points,vert,smp)] = np.nan # mask outside points ug = u.reshape((400,400)) # fold back into a grid # make a contour plot of the solution fig,ax = plt.subplots() p = ax.contourf(xg,yg,ug,cmap='viridis') ax.plot(nodes[:,0],nodes[:,1],'ko',markersize=4) for s in smp: ax.plot(vert[s,0],vert[s,1],'k-',lw=2) ax.set_aspect('equal') fig.colorbar(p,ax=ax) fig.tight_layout() plt.show()
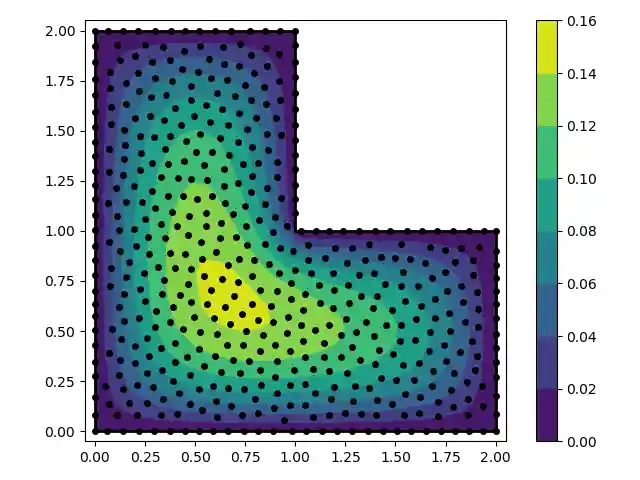
''' In this example we solve the Poisson equation over an L-shaped domain with fixed boundary conditions. We use the RBF-FD method. ''' import numpy as np from rbf.fd import weight_matrix from rbf.basis import phs3 from rbf.geometry import contains from rbf.nodes import menodes import matplotlib.pyplot as plt from scipy.sparse import vstack from scipy.sparse.linalg import spsolve from scipy.interpolate import LinearNDInterpolator # Define the problem domain with line segments. vert = np.array([[0.0,0.0],[2.0,0.0],[2.0,1.0], [1.0,1.0],[1.0,2.0],[0.0,2.0]]) smp = np.array([[0,1],[1,2],[2,3],[3,4],[4,5],[5,0]]) N = 500 # total number of nodes. n = 20 # stencil size. basis = phs3 # radial basis function used to compute the weights. order = 2 # Order of the added polynomials. # generate nodes nodes,smpid = menodes(N,vert,smp) edge_idx, = (smpid>=0).nonzero() interior_idx, = (smpid==-1).nonzero() # create "left hand side" matrix A_int = weight_matrix(nodes[interior_idx],nodes,diffs=[[2,0],[0,2]], n=n,basis=basis,order=order) A_edg = weight_matrix(nodes[edge_idx],nodes,diffs=[0,0]) A = vstack((A_int,A_edg)) # create "right hand side" vector d_int = -1*np.ones_like(interior_idx) d_edg = np.zeros_like(edge_idx) d = np.hstack((d_int,d_edg)) # find the solution at the nodes u_soln = spsolve(A,d) # interpolate the solution on a grid xg,yg = np.meshgrid(np.linspace(-0.05,2.05,400),np.linspace(-0.05,2.05,400)) points = np.array([xg.flatten(),yg.flatten()]).T u_itp = LinearNDInterpolator(nodes,u_soln)(points) # mask points outside of the domain u_itp[~contains(points,vert,smp)] = np.nan ug = u_itp.reshape((400,400)) # fold back into a grid # make a contour plot of the solution fig,ax = plt.subplots() p = ax.contourf(xg,yg,ug,cmap='viridis') ax.plot(nodes[:,0],nodes[:,1],'ko',markersize=4) for s in smp: ax.plot(vert[s,0],vert[s,1],'k-',lw=2) ax.set_aspect('equal') fig.colorbar(p,ax=ax) fig.tight_layout() plt.show()