Paired REad TEXTure Viewer. OpenGL Powered Pretext Contact Map Viewer.
PretextView is a desktop application for viewing pretext contact maps.
Usage
Home screen
Escormiddle mouse button: enter main GUI.right mouse button: holding down and move to drag the Hi-C figure move within the window.Mouse scrollto zoom. A three button mouse is recomended.E: enter the Edit mode.F: enter the Select sort area mode.M: enter the Meta Tag edit mode.S: enter the Scaffold painting mode.X: enter the extension edit mode.W: enter the Waypoint edit mode.J: window jump to the diagonal line without changing the zoom-level, which is useful for selecting the correct place for a small fragment.L: toggle the Grid Line.T: toggle the Tooltip. When it is on, the tooltip window shows the contig names and positions for the row and column under the cursor, the scaffold id, meta tag names that apply at that locus (from Meta Tag mode), and values for enabled extension tracks (e.g. coverage) when those are turned on.I: toggle the ID bar.3: toggle the 3p_telomere extension.5: toggle the 5p_telomere extension.Up/Down: change the color map.Left/Right: decrease / increase the Gamma mid (default: 0.5).
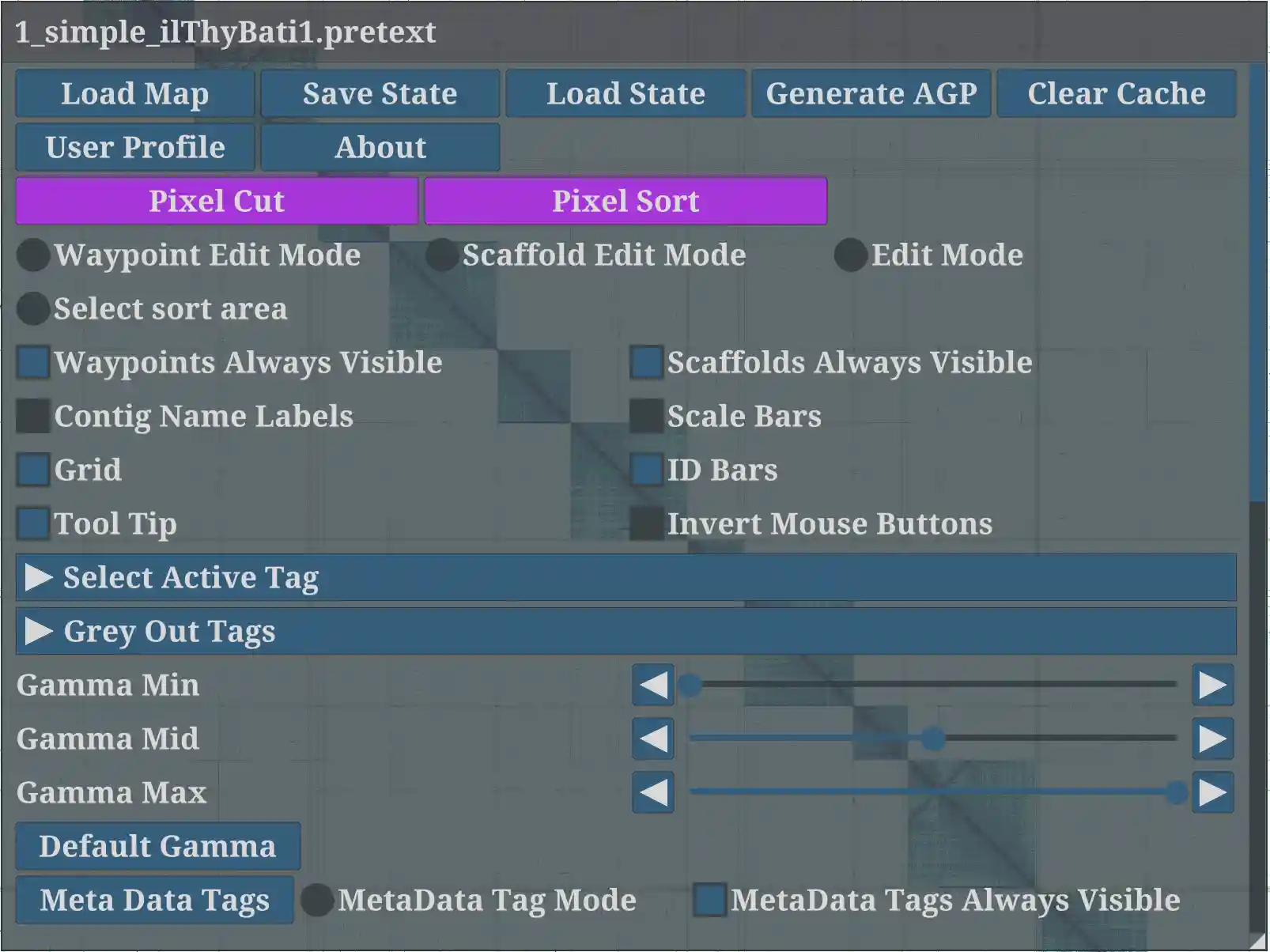
Main GUI (Esc)
Load Map: select.pretextfile to load the HiC figure.Save State: Save a specific to contain edits, painted scaffolds information, meta tags, waypoints, camera position, zoom level, window settings. Actually, the history is automatically saved into a file with hashed name in the cache file directory (for example,/Users/[yourUserName]/.config/PretextView/on mac).Load State: select a file to load the state.Generate AGP: save the curated genome into the.agpformat.Clear Cache: Clear all edits and revert to the pre-curation state. Please note this action is irreversible.Clipboard Editor: opens a text buffer tied to the system clipboard. Use Ctrl+C / Ctrl+V / Ctrl+X to copy, paste, and cut (Cmd+C / Cmd+V / Cmd+X on macOS). Save as opens a file dialog to save the buffer (default filename suffix_clipboard.txt).
Edit mode (E)
left mouse button: Click and drag with the left mouse button to select an are to zoom to.- Pickup a region of a contig with the left mouse button, pickup a whole contig with the middle mouse button or spacebar. Place a region with the left mouse button. Invert a selected region with the middle mouse button or spacebar. Undo the last edit with the 'q' key. Exit edit mode with the 'e' key. Use the GUI to see a list of completed edits.
P: copy the highlighted map range to the system clipboard. The text lists each map fragment in the selection with the contig name and a local span in megabases along that fragment (not a single genome-wide coordinate).V: break (split) the contig at the start of the current selection.
Waypoint mode (W)
left mouse button: place a waypoint.middle mouse buttonorspacebar: delete a waypoint.W: Exit waypoint mode.L: change the way point lines to vertical, horizontal or cross.- Use the GUI to see a list of waypoints, click on a waypoint in the list to vist it.
Scaffold edit mode (S)
Enter scaffolding mode with the 's' key.
- Use the GUI to see a list of scaffolds.
Extension edit mode (X)
Enter extension mode with the X key to toggle various genomic feature overlays. Once in extension mode, press:
3: Toggle 3p_telomere track on/off (also works at top level)5: Toggle 5p_telomere track on/off (also works at top level)T: Toggle general telomere track on/offC: Toggle coverage track on/offG: Toggle gap track on/offR: Toggle repeat_density track on/offX: Exit extension mode
Note: The 3 and 5 keys work both inside and outside Extension Mode. Other shortcuts (C, G, R, T) only work when in Extension Mode, as these keys have different functions outside of it.
Select sort area mode (F)
After click Pixel Sort button in the main UI, it will defaultly run sort globally. If enter the the select sort area mode(by pressing F), it can sort only the selected area with pressing Space after selecting at least 2 contigs.
Enter the select sort area mode by pressing F.
Left mouse click: select / un-select area for sorting.S: clear all the select area.Space: callPixel Sortto sort the select area (NOTE: only work if select at least 3 fragments).Q/W: quit/redo edit (currently will change the edit made globally not only the edit made within the selected area, so use this with caution as it can change also other parts)C: cut the scaffolds/contigs within the selected area.Z: glue the splitted fragments within the selected area. (NOTE: only link the continuous fragments.)
Sort fragments according to link score
Usage, first endter the main GUI:
-
Right click the
Pixel Sortbutton to open sort settings,-
Smallest Frag Size (pixels)(defaul: 2) which represents the smallest fragment size in unit of pixels to consider during the sorting -
Link Score Threshold: $S_{thresh}$ (defaul: 0.4). - Select the sort mode:
-
UnionFind: sort all links higher than $S_{thresh}$ and then traverse all the links by from high to low. -
Fuse: traverse all links and fuse two chains.
-
-
-
Left click the
Pixel Sortbutton: run. - Redo all edits
- Erase all edits
Pixel Cut according to the continuity of HiC density
Usage: In the main GUI:
- Right click the
Pixel Cutbutton to open cut settings,Cut threshold: the smaller, the more likely to break one contig at the discontinuous locations.Pixel_mean window size: length of range from diagonl line, which is considered to calculate the HiC density.Smallest frag size: the smallest fragment length that can be generated.
- Left click the
Pixel Cutbutton: cut all the scaffolds/contigs globally.
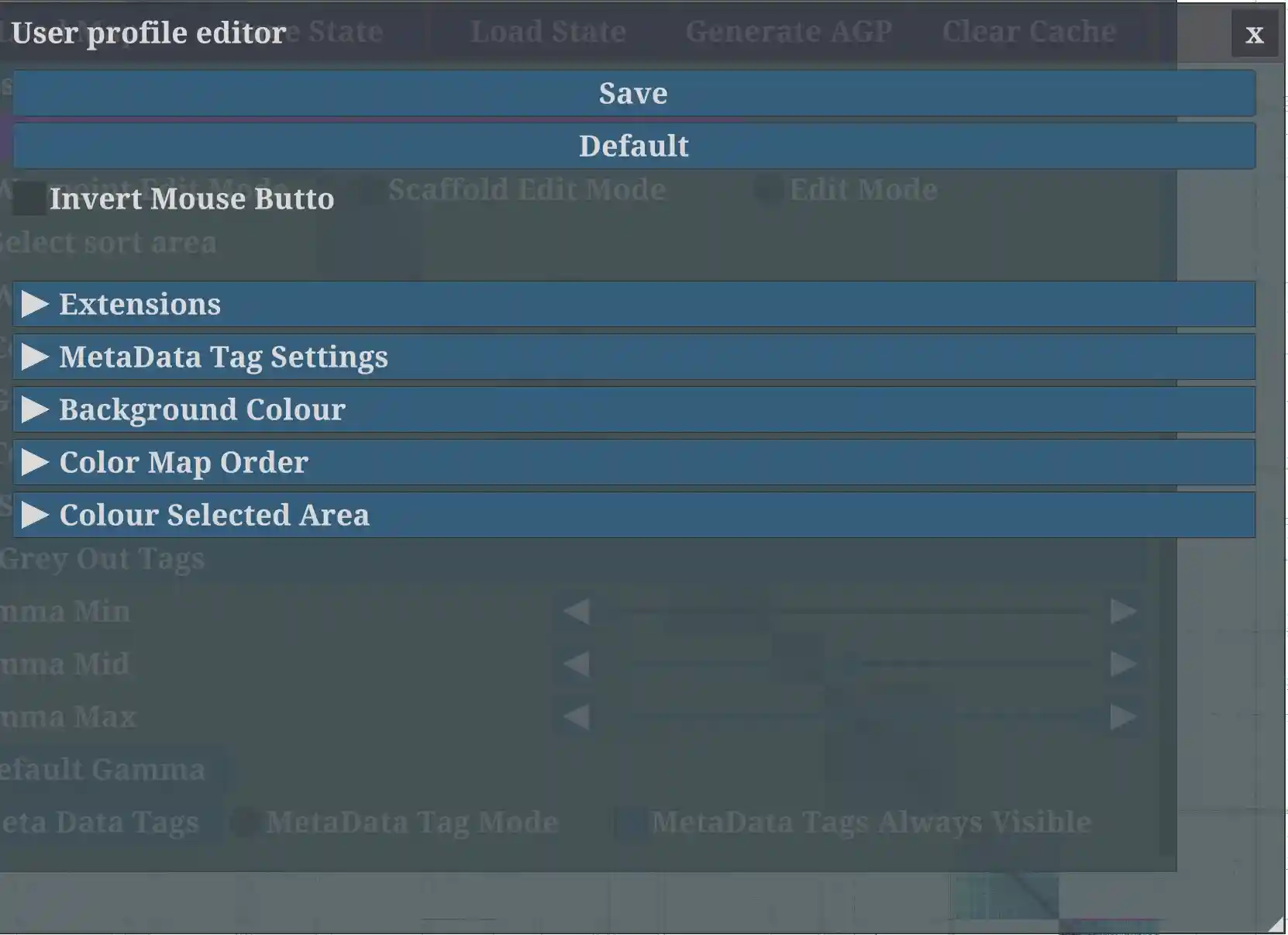
User Profile
Click the User Profile button in main GUI.
Set the Meta tags, Background Color, Color Map(HiC figure), Extension settings and the Colour Selected Area.
Saving
Map state is automatically saved ($XDG_CONFIG_DIR or ~/.config on Unix, and the %APPDATA% folder on Windows) while the app runs, and is loaded on map load.
You can also manually save/load state via the UI.
AGP Output
Map state can be output in AGP format via the UI. Objects are first created according to the scaffolds defined in scaffolding mode, with remaining sequences output as singletons.
AGP Correction
Note that object/part sizes will only be accurate up to the size of an individual map texel, and that any input sequences smaller than an individual texel will not be output.
AGP files can be corrected by the included python script AGPCorrect, which requires access to the input sequences in (gzipped) FASTA format.
AGPCorrect /path/to/ref.fa(.gz) /path/to/current/map.agp >/path/to/corrected_scaffs.agpThe script requires
- Python >= 3.8
- Biopython
Requirments, running
OpenGL 3.3
2G of RAM
Windows, Mac and Linux Builds
The prebuilt apps for Windows, Mac and Linux are available here.
The Mac app was built on MacOS 10.13.6
The Linux app was built on kernel 3.13
The Windows app was build on Windows 10, and is known to work on at least Windows 7.
Third-Party acknowledgements
PretextView uses the following third-party libraries:
- glfw
- libdeflate
- FontStash
- Nuklear
- glad
- stb_image
- stb_image_write
- stb_truetype
- stb_sprintf
- Fonts from the 'Droid Serif' font set
- Icons from open-iconic
- fmt
Installation
Requires:
- clang >= 11.0.0 [Unix]
- clang-cl >= 11.0.0 [Windows]
- cmake >= 3.1.7
# git submodule update --init --recursive # Adding dependencies from third-party libraries which is contained in the install script ./install.cmake.sh # [Unix] install.cmake.bat # [Windows]
Application will be installed to the 'app' folder in the source directory.
NOTE for Mac user who downloads this from Release page. If the user is told that the software is damaged, please unzip the file and run
xattr -d com.apple.quarantine /path/to/the/downloaded/dmg/file
to remove the quarantine and then unmount and re-mount the dmg file. And then try to open the PretextViewAI.app.
Details of sorting algorithm
Link score calculation
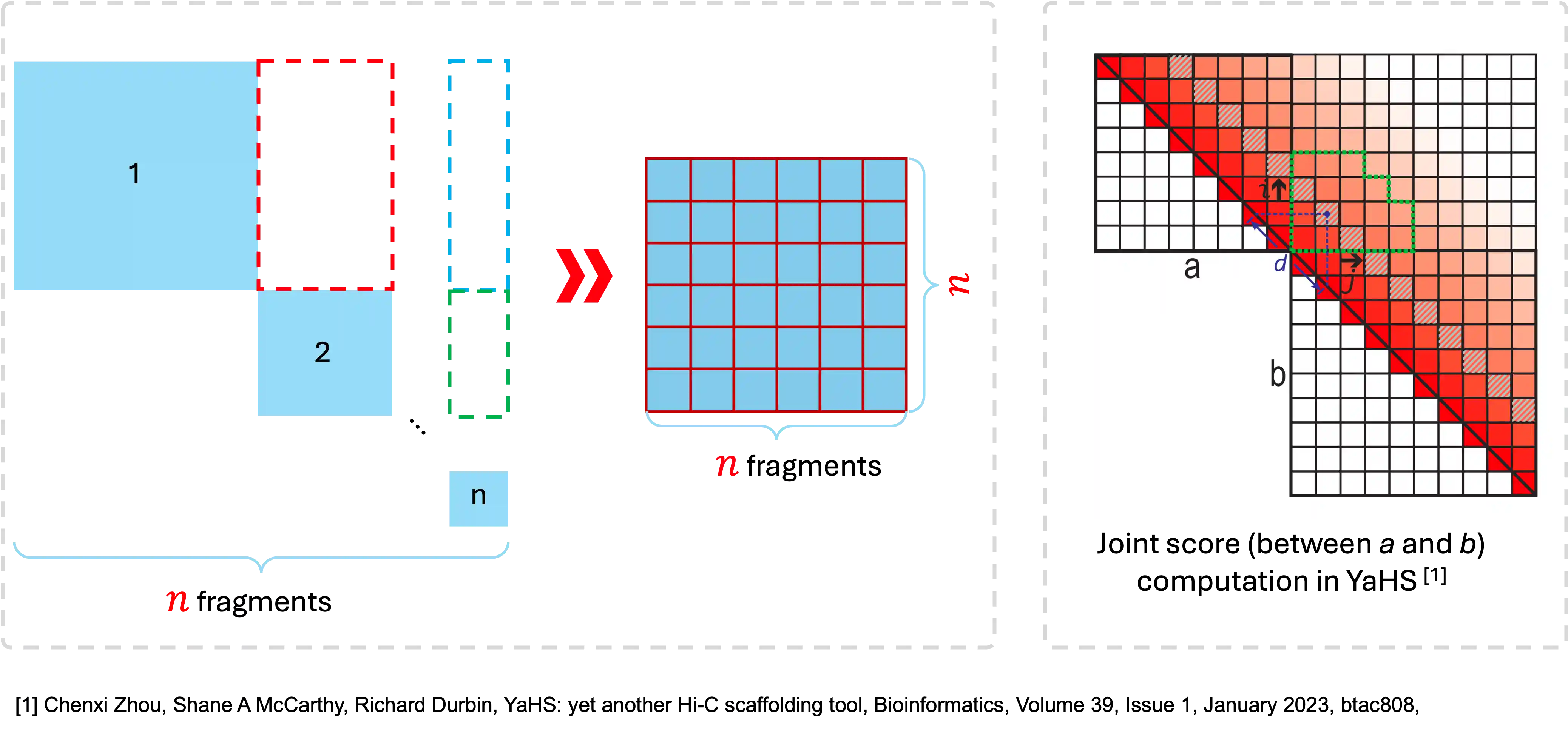
The score between two fragments represents the level of continuity. The higher score the better continuity.
The link score calculation method refers to YaHS.
Reference: Chenxi Zhou, Shane A McCarthy, Richard Durbin, YaHS: yet another Hi-C scaffolding tool, Bioinformatics, Volume 39, Issue 1, January 2023, btac808, https://doi.org/10.1093/bioinformatics/btac808
Union Find Mode
- Select and sort all links with a score higher than the threshold (default: 0.4).
- Traverse all the links from highest to lowest score, consider the the two ends.
Suppose we have two chains: $$A: [a_1, a_2, a_3, \ldots, a_i, \ldots, a_n]$$ $$B: [b_1, b_2, b_3, \ldots, b_j, \ldots, b_m]$$ They can only be linked if there is a strong (larger than threshold) link between one of the following pairs:$[a_1, b_1], [a_1, b_m], [a_n, b_1], [a_n, b_m]$
Fuse Mode & Deep Fuse
For two chains: $A: [a_1, a_2, a_3, \ldots, a_i, \ldots, a_n]$ and $B: [b_1, b_2, b_3, \ldots, b_j, \ldots, b_m]$
The Union Find approach considers only the two ends. Now we consider fusing these two chains into one if $[a_i, b_j]$ is a link with a score higher than the threshold.
In fuse mode, the tail of $A$, denoted $A_t$: $[a_i, a_{i+1}, \ldots, a_n]$ and the head of $B$, denoted $B_h$: $[b_1, b_2, \ldots, b_j]$ are fused. Let $a_{i-1}$ be the source, and $b_{j+1}$ be the sink. We insert $A_t$ and $B_h$ in the optimal way to achieve the highest total score, preserving their internal order.
If the link score of the fused chain $S_F$ is larger than the original scores plus threshold: $$S_F = \sum_{i=0}^{N-1} s(f_i, f_{i+1}) \ge \Bigl(S_{A_t} + S_{B_h} + S_{\text{thresh}}\Bigr)$$ where $f \in F$, and $F$ is the chain fused from $A_t$ and $B_h$. The score of $A_t$ is $S_{A_t} = \sum_{i=0}^{N_{A_t}-1} s(a_i, a_{i+1})$ with $a \in A_t$.
Fuse mode:
Runs the fuse process after Union Find. Suitable for samples with fewer than 800 fragments.
Deep fuse:
Runs the fuse process without using Union Find, suitable for samples with fewer than 100 fragments.