Using LSTM autoencoder, L1 Regularization
Purpose
- For anomaly detection, autoencoder is widely used.
- But using autoencoder, which have many variables with strong correlations, is said to cause a decline of detection power.
- To avoid the above problem, the technique to apply L1 regularization to LSTM autoencoder is advocated in the below paper.
N. Gugulothu, P. Malhotra, L. Vig, and G. Shroff, “Sparse neural networks for anomaly detection in high-dimensional time series,” in AI4IOT Workshop in Conjunction with ICML, International Joint Conference on Artificial Intelligence and European Conference on Artificial Intelligence, Stockholm, Sweden, 2018.
- The point is to use L1 regularization at the second layer of the sequence model(right under the input data).
Algorithm and How to implement
- For the implementation, tensorflow and keras are used.
Structure of layers
def create_dataset(self, X, y, time_steps=1): self.X = X self.y = y self.Xs, self.ys = [], [] self.time_steps = time_steps for self.i in range(len(self.X) - self.time_steps): self.v = self.X.iloc[self.i:(self.i + self.time_steps)].values self.Xs.append(self.v) self.ys.append(self.y.iloc[self.i + self.time_steps]) return np.array(self.Xs), np.array(self.ys)
-
At first, define "Standard RNN EncoderDecoder", then define "Sparse RNN Encoder-Decoder".
-
Sparse RNN Encoder-Decoder is built by adding some changes to Standard RNN EncoderDecoder as below;
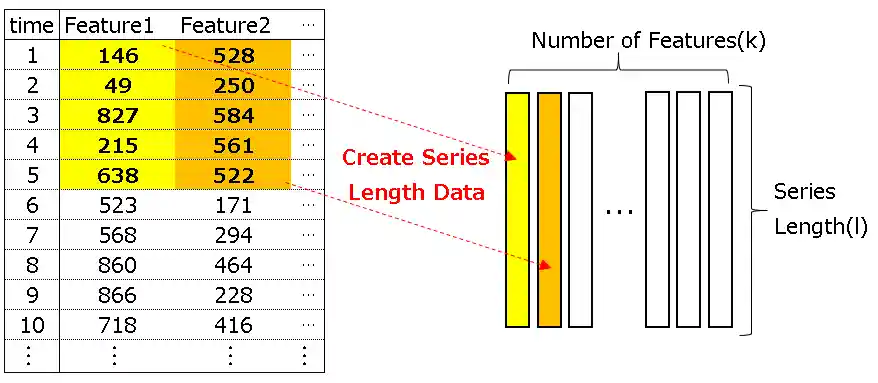
- Flatten the input
- Insert the custom layer(with L1 regularization)
- Reshape the 2. output
-
Structure of Standard RNN EncoderDecoder
def Usual_LSTM(X): hidden = 5 timesteps=X.shape[1] num_features=X.shape[2] model = Sequential([ LSTM(hidden, input_shape=(timesteps, num_features)), RepeatVector(timesteps), LSTM(hidden, return_sequences=True), TimeDistributed(Dense(num_features)) ]) model.summary() model.compile(loss='mse', optimizer='adam') return model
| Seq | Layer | Input Shape | Output Shape |
|---|---|---|---|
| 1 | LSTM | (None, l, k) | (None, h) |
| 2 | RepeatVector | (None, h) | (None, l, h) |
| 3 | LSTM | (None, l, h) | (None, l, h) |
| 4 | TimeDistributed | (None, l, h) | (None, l ,k) |
- Structure of Sparse RNN Encoder-Decoder
class MyLayer(tf.keras.layers.Layer): def __init__(self, units): super().__init__() self.units = units def build(self, input_shape): self.kernel = self.add_weight( "kernel", shape=[1, self.units], initializer='uniform', trainable=True, regularizer = tf.keras.regularizers.l1(0.001) ) def call(self, input): output = input*self.kernel return tf.nn.relu(output) def Sparse_LSTM(X): hidden = 5 timesteps=X.shape[1] num_features=X.shape[2] model = Sequential([ Flatten(input_shape=(timesteps, num_features)), MyLayer(timesteps*num_features), Reshape(target_shape=(timesteps, num_features)), LSTM(hidden, input_shape=(timesteps, num_features)), RepeatVector(timesteps), LSTM(hidden, return_sequences=True), TimeDistributed(Dense(num_features)) ]) model.compile( loss="mse", optimizer='adam') model.summary() return model
| Seq | Layer | Input Shape | Output Shape |
|---|---|---|---|
| 1 | Flatten | (None, l, k) | (None, l×k) |
| 2 | Custom | (None, l×k) | (None, l×k) |
| 3 | Reshape | (None, l×k) | (None, l, k) |
| 4 | LSTM | (None, l, k) | (None, h) |
| 5 | RepeatVector | (None, h) | (None, l, h) |
| 6 | LSTM | (None, l, h) | (None, l, h) |
| 7 | TimeDistributed | (None, l, h) | (None, l ,k) |
Results
- Assume that there are two types of data, the one is sine wave with noise(F1), and the other is normal random number with noise(F2).
- Assume two case, (1)[F1, F2](features are both independent), (2)[F1, F2, F2, F2, F2](partical Features are not independent)
-
Create train data with former 10,000 records, and define latter 5,000 records as test data.
-
How to plot Learning history
def plotting_history(self, history): self.history = history plt.figure(figsize=(4, 4)) plt.plot(self.history.history['loss'], label='Training Loss') plt.plot(self.history.history['val_loss'], label='Validation Loss') plt.xlabel("Epochs") plt.ylabel("loss") plt.legend() plt.show()
- How to compute Mahalanobis Distance
def compute_mahalanobis(model, X_train, X_test): train_error = model.predict(X_train) - X_train cov = np.cov(train_error.reshape(-1, X_train.shape[-1]).T) mean = np.mean(train_error.reshape(-1, X_train.shape[-1]), axis=0) test_error = model.predict(X_test) - X_test temp_reshape = test_error.reshape(-1, test_error.shape[-1]) return np.mean(np.array([distance.mahalanobis(mean, temp_reshape[i], cov) for i in range(len(temp_reshape))]).reshape(-1, X_train.shape[1]), axis=1)
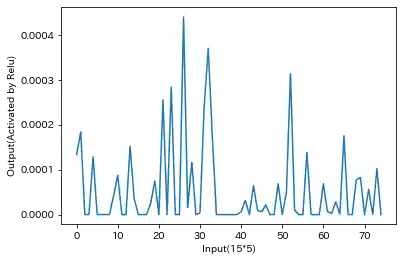
- Relu Output of Sparse RNN(Sample)
partial input equal to zero
Standard RNN
Case(1)
| Learning history | Mahalanobis Distance (=Anomaly Index) |
|---|---|
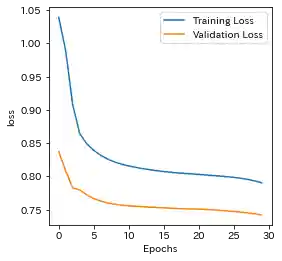 |
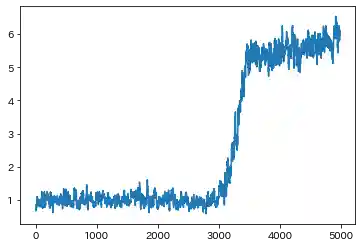 |
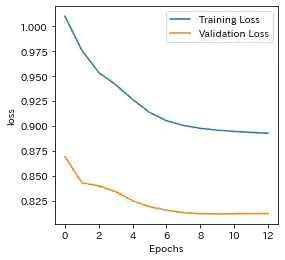
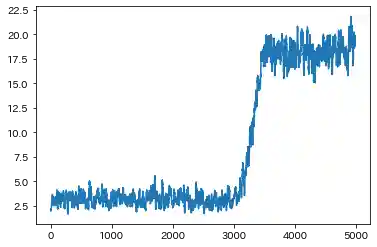
Case(2)
| Learning history | Mahalanobis Distance (=Anomaly Index) |
|---|---|
 |
 |
Sparse RNN
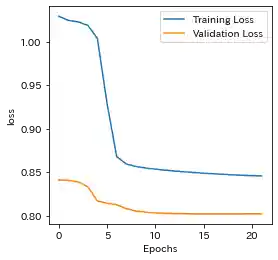
Case(1)
| Learning history | Mahalanobis Distance (=Anomaly Index) |
|---|---|
 |
 |
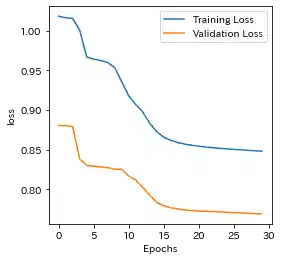
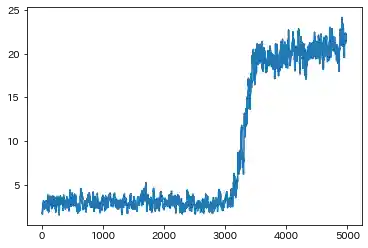
Case(2)
| Learning history | Mahalanobis Distance (=Anomaly Index) |
|---|---|
 |
 |
Conclustion
- Both results are almost the same. But in Case(2), Standard RNN could not capture the F1 ascending trend(seen in Case(1)), which is caused by sine wave.
- Sparse RNN seems to be able to capture the above trend, so it might have the ability to eliminate the effect of strong correlation among features to some extent.
- Sparse RNN seems that it could not learn well at first, so it should be noted that the patience of EarlyStopping has to be set as a somewhat higher number.