Smooth and Strongly Convex Nearest Brenier Potentials — POT Python Optimal Transport 0.9.6 documentation
Note
Go to the end to download the full example code.
Note
Example added in release: 0.9.2.
This example is designed to show how to use SSNB [58] in POT. SSNB computes an l-strongly convex potential \(\varphi\) with an L-Lipschitz gradient such that \(\nabla \varphi \# \mu \approx \nu\). This regularity can be enforced only on the components of a partition of the ambient space, which is a relaxation compared to imposing global regularity.
In this example, we consider a source measure \(\mu_s\) which is the uniform measure on the unit square in \(\mathbb{R}^2\), and the target measure \(\mu_t\) which is the image of \(\mu_x\) by \(T(x_1, x_2) = (x_1 + 2\mathrm{sign}(x_2), 2 * x_2)\). The map \(T\) is non-smooth, and we wish to approximate it using a “Brenier-style” map \(\nabla \varphi\) which is regular on the partition \(\lbrace x_1 <=0, x_1>0\rbrace\), which is well adapted to this particular dataset.
We represent the gradients of the “bounding potentials” \(\varphi_l, \varphi_u\) (from [59], Theorem 3.14), which bound any SSNB potential which is optimal in the sense of [58], Definition 1:
\[\varphi \in \mathrm{argmin}_{\varphi \in \mathcal{F}}\ \mathrm{W}_2(\nabla \varphi \#\mu_s, \mu_t),\]
where \(\mathcal{F}\) is the space functions that are on every set \(E_k\) l-strongly convex with an L-Lipschitz gradient, given \((E_k)_{k \in [K]}\) a partition of the ambient source space.
We perform the optimisation on a low amount of fitting samples and with few iterations, since solving the SSNB problem is quite computationally expensive.
THIS EXAMPLE REQUIRES CVXPY
# Author: Eloi Tanguy <eloi.tanguy@math.cnrs.fr> # License: MIT License # sphinx_gallery_thumbnail_number = 3 import matplotlib.pyplot as plt import numpy as np import ot
Generating the fitting data
n_fitting_samples = 30 rng = np.random.RandomState(seed=0) Xs = rng.uniform(-1, 1, size=(n_fitting_samples, 2)) Xs_classes = (Xs[:, 0] < 0).astype(int) Xt = np.stack([Xs[:, 0] + 2 * np.sign(Xs[:, 0]), 2 * Xs[:, 1]], axis=-1) plt.scatter( Xs[Xs_classes == 0, 0], Xs[Xs_classes == 0, 1], c="blue", label="source class 0" ) plt.scatter( Xs[Xs_classes == 1, 0], Xs[Xs_classes == 1, 1], c="dodgerblue", label="source class 1", ) plt.scatter(Xt[:, 0], Xt[:, 1], c="red", label="target") plt.axis("equal") plt.title("Splitting sphere dataset") plt.legend(loc="upper right") plt.show()
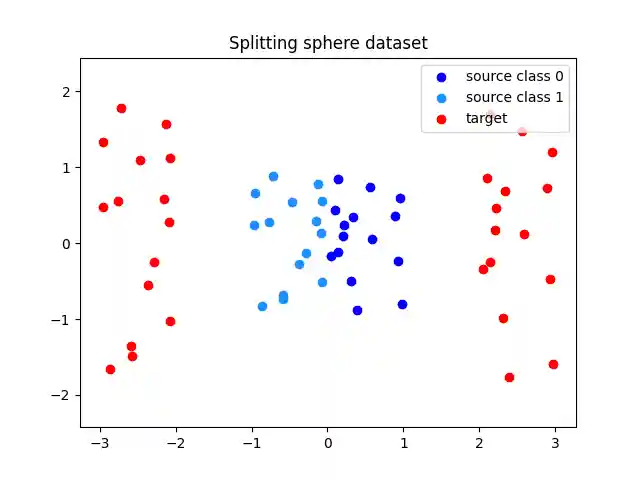
Fitting the Nearest Brenier Potential
Plotting the images of the source data
plt.clf() plt.scatter(Xs[:, 0], Xs[:, 1], c="dodgerblue", label="source") plt.scatter(Xt[:, 0], Xt[:, 1], c="red", label="target") for i in range(n_fitting_samples): plt.plot([Xs[i, 0], G[i, 0]], [Xs[i, 1], G[i, 1]], color="black", alpha=0.5) plt.title("Images of in-data source samples by the fitted SSNB") plt.legend(loc="upper right") plt.axis("equal") plt.show()
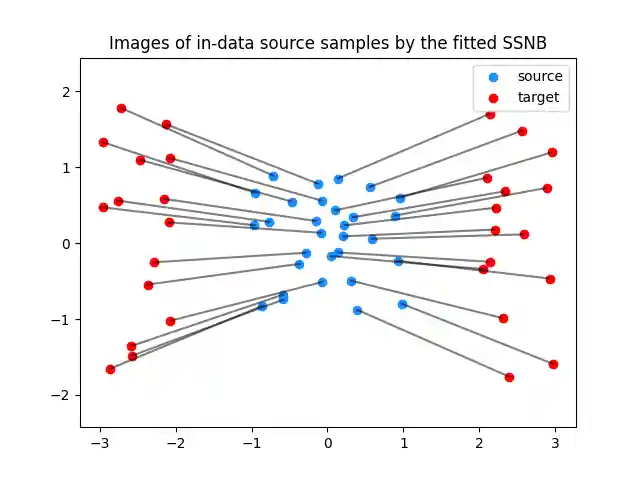
Computing the predictions (images by nabla phi) for random samples of the source distribution
n_predict_samples = 50 Ys = rng.uniform(-1, 1, size=(n_predict_samples, 2)) Ys_classes = (Ys[:, 0] < 0).astype(int) phi_lu, G_lu = ot.mapping.nearest_brenier_potential_predict_bounds( Xs, phi, G, Ys, Xs_classes, Ys_classes, gradient_lipschitz_constant=L )
Plot predictions for the gradient of the lower-bounding potential
plt.clf() plt.scatter(Xs[:, 0], Xs[:, 1], c="dodgerblue", label="source") plt.scatter(Xt[:, 0], Xt[:, 1], c="red", label="target") for i in range(n_predict_samples): plt.plot( [Ys[i, 0], G_lu[0, i, 0]], [Ys[i, 1], G_lu[0, i, 1]], color="black", alpha=0.5 ) plt.title("Images of new source samples by $\\nabla \\varphi_l$") plt.legend(loc="upper right") plt.axis("equal") plt.show()
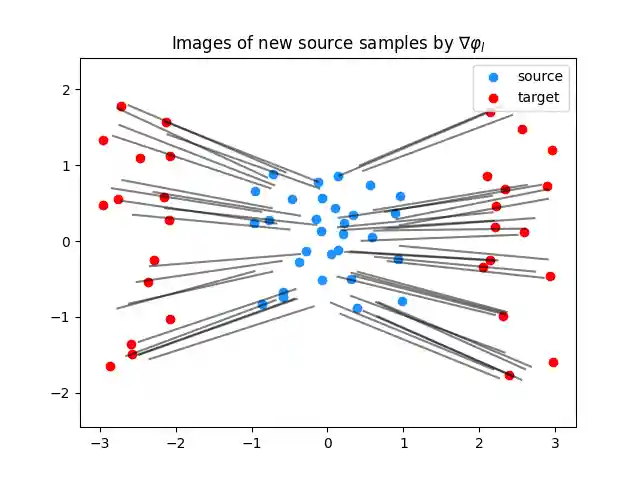
Plot predictions for the gradient of the upper-bounding potential
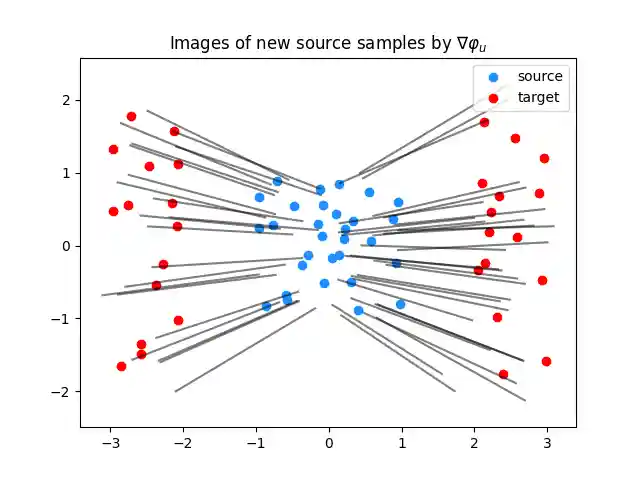
plt.clf() plt.scatter(Xs[:, 0], Xs[:, 1], c="dodgerblue", label="source") plt.scatter(Xt[:, 0], Xt[:, 1], c="red", label="target") for i in range(n_predict_samples): plt.plot( [Ys[i, 0], G_lu[1, i, 0]], [Ys[i, 1], G_lu[1, i, 1]], color="black", alpha=0.5 ) plt.title("Images of new source samples by $\\nabla \\varphi_u$") plt.legend(loc="upper right") plt.axis("equal") plt.show()
Total running time of the script: (0 minutes 57.478 seconds)