1D Wasserstein barycenter demo for Unbalanced distributions — POT Python Optimal Transport 0.9.6 documentation
Note
Go to the end to download the full example code.
This example illustrates the computation of regularized Wasserstein Barycenter as proposed in [10] for Unbalanced inputs.
[10] Chizat, L., Peyré, G., Schmitzer, B., & Vialard, F. X. (2016). Scaling algorithms for unbalanced transport problems. arXiv preprint arXiv:1607.05816.
# Author: Hicham Janati <hicham.janati@inria.fr> # # License: MIT License # sphinx_gallery_thumbnail_number = 4 import numpy as np import matplotlib.pylab as pl import ot # necessary for 3d plot even if not used from mpl_toolkits.mplot3d import Axes3D # noqa from matplotlib.collections import PolyCollection
Generate data
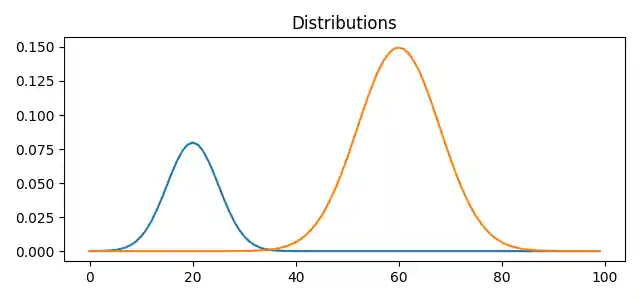
Plot data
# plot the distributions pl.figure(1, figsize=(6.4, 3)) for i in range(n_distributions): pl.plot(x, A[:, i]) pl.title("Distributions") pl.tight_layout()
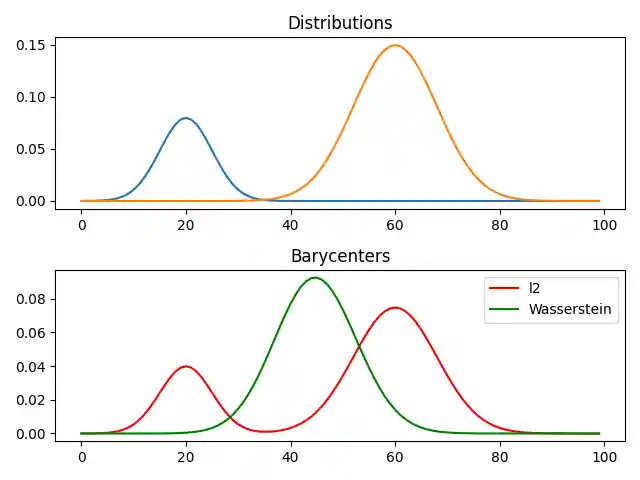
Barycenter computation
# non weighted barycenter computation weight = 0.5 # 0<=weight<=1 weights = np.array([1 - weight, weight]) # l2bary bary_l2 = A.dot(weights) # wasserstein reg = 1e-3 alpha = 1.0 bary_wass = ot.unbalanced.barycenter_unbalanced(A, M, reg, alpha, weights=weights) pl.figure(2) pl.clf() pl.subplot(2, 1, 1) for i in range(n_distributions): pl.plot(x, A[:, i]) pl.title("Distributions") pl.subplot(2, 1, 2) pl.plot(x, bary_l2, "r", label="l2") pl.plot(x, bary_wass, "g", label="Wasserstein") pl.legend() pl.title("Barycenters") pl.tight_layout()
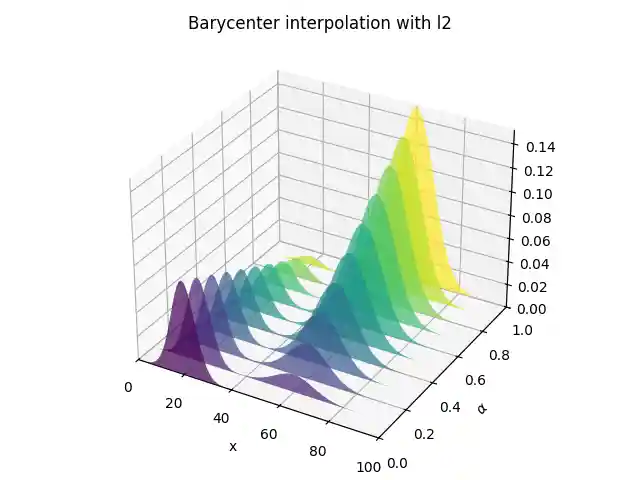
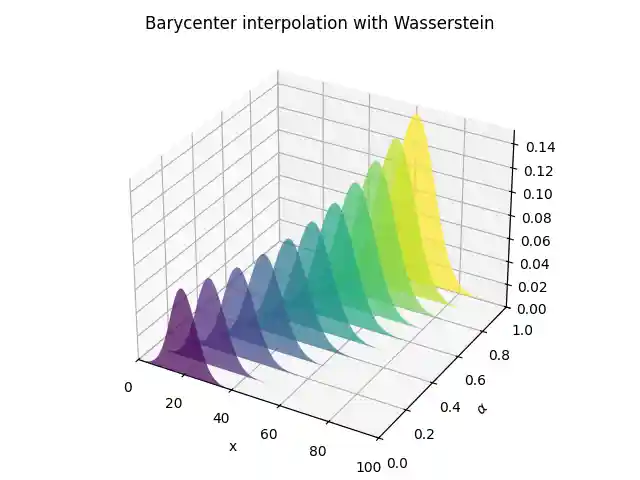
Barycentric interpolation
# barycenter interpolation n_weight = 11 weight_list = np.linspace(0, 1, n_weight) B_l2 = np.zeros((n, n_weight)) B_wass = np.copy(B_l2) for i in range(0, n_weight): weight = weight_list[i] weights = np.array([1 - weight, weight]) B_l2[:, i] = A.dot(weights) B_wass[:, i] = ot.unbalanced.barycenter_unbalanced( A, M, reg, alpha, weights=weights ) # plot interpolation pl.figure(3) cmap = pl.get_cmap("viridis") verts = [] zs = weight_list for i, z in enumerate(zs): ys = B_l2[:, i] verts.append(list(zip(x, ys))) ax = pl.gcf().add_subplot(projection="3d") poly = PolyCollection(verts, facecolors=[cmap(a) for a in weight_list]) poly.set_alpha(0.7) ax.add_collection3d(poly, zs=zs, zdir="y") ax.set_xlabel("x") ax.set_xlim3d(0, n) ax.set_ylabel(r"$\alpha$") ax.set_ylim3d(0, 1) ax.set_zlabel("") ax.set_zlim3d(0, B_l2.max() * 1.01) pl.title("Barycenter interpolation with l2") pl.tight_layout() pl.figure(4) cmap = pl.get_cmap("viridis") verts = [] zs = weight_list for i, z in enumerate(zs): ys = B_wass[:, i] verts.append(list(zip(x, ys))) ax = pl.gcf().add_subplot(projection="3d") poly = PolyCollection(verts, facecolors=[cmap(a) for a in weight_list]) poly.set_alpha(0.7) ax.add_collection3d(poly, zs=zs, zdir="y") ax.set_xlabel("x") ax.set_xlim3d(0, n) ax.set_ylabel(r"$\alpha$") ax.set_ylim3d(0, 1) ax.set_zlabel("") ax.set_zlim3d(0, B_l2.max() * 1.01) pl.title("Barycenter interpolation with Wasserstein") pl.tight_layout() pl.show()
/home/circleci/project/ot/unbalanced/_sinkhorn.py:1666: RuntimeWarning: overflow encountered in divide
u = (A / Kv) ** fi
/home/circleci/project/ot/unbalanced/_sinkhorn.py:1671: RuntimeWarning: invalid value encountered in divide
v = (Q / Ktu) ** fi
/home/circleci/project/ot/unbalanced/_sinkhorn.py:1682: UserWarning: Numerical errors at iteration 595
warnings.warn("Numerical errors at iteration %s" % i)
/home/circleci/project/ot/unbalanced/_sinkhorn.py:1671: RuntimeWarning: overflow encountered in divide
v = (Q / Ktu) ** fi
/home/circleci/project/ot/unbalanced/_sinkhorn.py:1682: UserWarning: Numerical errors at iteration 974
warnings.warn("Numerical errors at iteration %s" % i)
/home/circleci/project/ot/unbalanced/_sinkhorn.py:1682: UserWarning: Numerical errors at iteration 615
warnings.warn("Numerical errors at iteration %s" % i)
/home/circleci/project/ot/unbalanced/_sinkhorn.py:1682: UserWarning: Numerical errors at iteration 455
warnings.warn("Numerical errors at iteration %s" % i)
/home/circleci/project/ot/unbalanced/_sinkhorn.py:1682: UserWarning: Numerical errors at iteration 361
warnings.warn("Numerical errors at iteration %s" % i)
Total running time of the script: (0 minutes 1.258 seconds)