Using the PASTE algorithm — paste3 src documentation
# This cell is to allow automatic notebook generation for docs # You may want to comment this out if you have paste3 installed import sys from pathlib import Path sys.path.insert(0, str(Path.cwd().parent.parent.parent / "src"))
This noteook highlights the creation of slices (Anndata objects), usage of the pairwise_align and center_align functions of paste3, along with stacking and plotting functionalities.
This notebook primarily highlights how you would use the ``paste3`` package in ``PASTE`` (i.e. full alignment) mode, when the slices overlap over the full 2D assayed region, with a similar field of view and similar number and proportion of cell types.
import time import matplotlib.patches as mpatches import matplotlib.pyplot as plt import numpy as np import pandas as pd import scanpy as sc import torch from paste3.helper import get_common_genes, match_spots_using_spatial_heuristic from paste3.paste import center_align, pairwise_align from paste3.visualization import plot_slice, stack_slices_center, stack_slices_pairwise
Read data and create AnnData objects
data_dir = "../../../tests/data/" # Assume that the coordinates of slices are named slice_name + "_coor.csv" def load_slices(data_dir, slice_names): slices = [] for slice_name in slice_names: slice_i = sc.read_csv(data_dir + slice_name + ".csv") slice_i_coor = np.genfromtxt(data_dir + slice_name + "_coor.csv", delimiter=",") slice_i.obsm["spatial"] = slice_i_coor # Preprocess slices sc.pp.filter_genes(slice_i, min_counts=15) sc.pp.filter_cells(slice_i, min_counts=100) slices.append(slice_i) return slices slices = load_slices(data_dir, ["slice1", "slice2", "slice3", "slice4"]) slice1, slice2, slice3, slice4 = slices
Each AnnData object consists of a gene expression matrx and spatial coordinate matrix.
array([[12., 0., 6., ..., 0., 0., 0.],
[ 7., 0., 1., ..., 1., 0., 0.],
[15., 1., 4., ..., 0., 0., 1.],
...,
[ 0., 0., 0., ..., 0., 0., 0.],
[ 1., 0., 0., ..., 0., 0., 0.],
[ 5., 0., 1., ..., 1., 0., 0.]], dtype=float32)
slice1.obsm["spatial"][0:5, :]
array([[13.064, 6.086],
[12.116, 7.015],
[13.945, 6.999],
[12.987, 7.011],
[15.011, 7.984]])
Note, you can choose to label the spots however you want. In this case, we use the default coordinates.
| n_counts | |
|---|---|
| 13.064x6.086 | 2181.0 |
| 12.116x7.015 | 2295.0 |
| 13.945x6.999 | 3375.0 |
| 12.987x7.011 | 2935.0 |
| 15.011x7.984 | 2964.0 |
| ... | ... |
| 21.953x24.847 | 541.0 |
| 20.98x24.963 | 860.0 |
| 20.063x24.964 | 508.0 |
| 19.007x25.045 | 626.0 |
| 21.957x25.871 | 2515.0 |
254 rows × 1 columns
| n_counts | |
|---|---|
| GAPDH | 2233.0 |
| UBE2G2 | 78.0 |
| MAPKAPK2 | 255.0 |
| NDUFA7 | 96.0 |
| ASNA1 | 172.0 |
| ... | ... |
| DIP2C | 31.0 |
| LYPLA2 | 19.0 |
| RGP1 | 24.0 |
| BPGM | 17.0 |
| HPS6 | 16.0 |
7998 rows × 1 columns
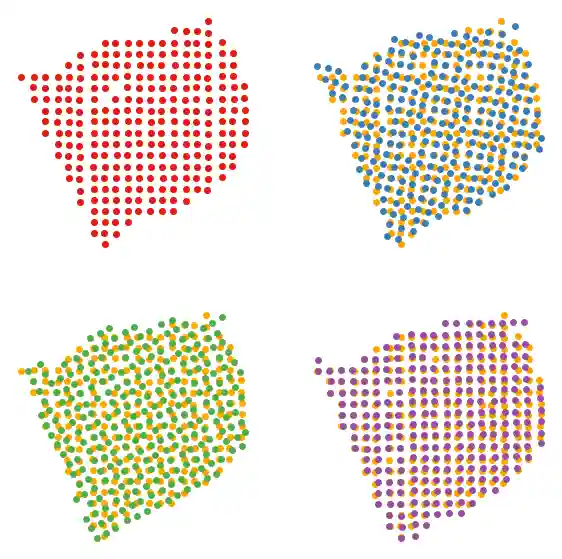
We can visualize the spatial coordinates of our slices using plot_slices.
slice_colors = ["#e41a1c", "#377eb8", "#4daf4a", "#984ea3"] fig, axs = plt.subplots(2, 2, figsize=(7, 7)) plot_slice(slice1, slice_colors[0], ax=axs[0, 0]) plot_slice(slice2, slice_colors[1], ax=axs[0, 1]) plot_slice(slice3, slice_colors[2], ax=axs[1, 0]) plot_slice(slice4, slice_colors[3], ax=axs[1, 1]) plt.show()
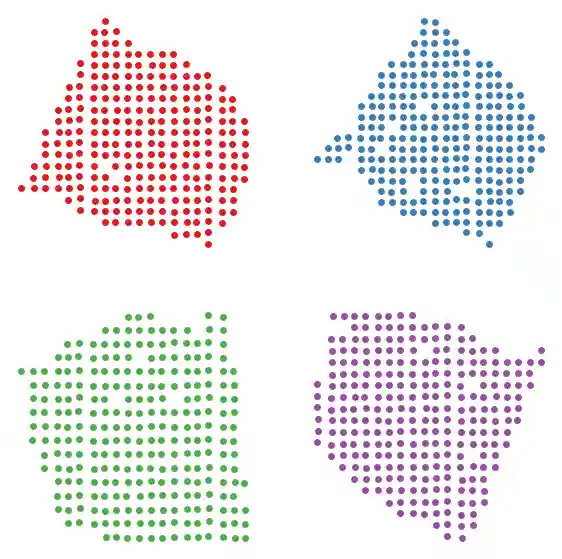

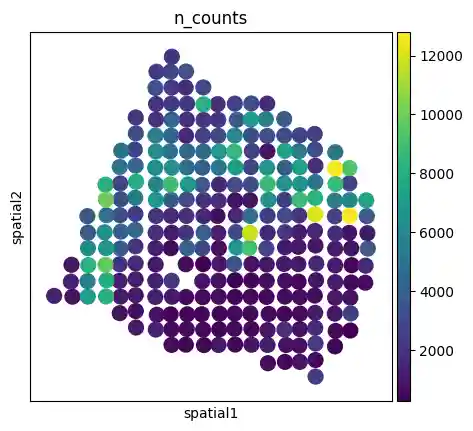
We can also plot using Scanpy’s spatial plotting function.
sc.pl.spatial(slice1, color="n_counts", spot_size=1)
Pairwise Alignment
Run PASTE pairwise_align.
start = time.time() pi12, _ = pairwise_align(slice1, slice2) pi23, _ = pairwise_align(slice2, slice3) pi34, _ = pairwise_align(slice3, slice4) print("Runtime: " + str(time.time() - start))
(INFO) (paste.py) (10-Jan-25 17:16:43) GPU is not available, resorting to torch CPU. (INFO) (paste.py) (10-Jan-25 17:16:43) GPU is not available, resorting to torch CPU. (INFO) (paste.py) (10-Jan-25 17:16:44) GPU is not available, resorting to torch CPU. Runtime: 0.3711864948272705
pd.DataFrame(pi12.cpu().numpy())
| 0 | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | ... | 240 | 241 | 242 | 243 | 244 | 245 | 246 | 247 | 248 | 249 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 0.003937 | 0.000000 | 0.000000 | 0.0 | 0.0 | 0.000000 | 0.000000 | 0.0 | 0.0 | 0.0 | ... | 0.0 | 0.000000 | 0.000000 | 0.000000 | 0.0 | 0.0 | 0.0 | 0.000000e+00 | 0.000000 | 0.000000 |
| 1 | 0.000063 | 0.003874 | 0.000000 | 0.0 | 0.0 | 0.000000 | 0.000000 | 0.0 | 0.0 | 0.0 | ... | 0.0 | 0.000000 | 0.000000 | 0.000000 | 0.0 | 0.0 | 0.0 | 0.000000e+00 | 0.000000 | 0.000000 |
| 2 | 0.000000 | 0.000000 | 0.000000 | 0.0 | 0.0 | 0.003937 | 0.000000 | 0.0 | 0.0 | 0.0 | ... | 0.0 | 0.000000 | 0.000000 | 0.000000 | 0.0 | 0.0 | 0.0 | 0.000000e+00 | 0.000000 | 0.000000 |
| 3 | 0.000000 | 0.000126 | 0.003811 | 0.0 | 0.0 | 0.000000 | 0.000000 | 0.0 | 0.0 | 0.0 | ... | 0.0 | 0.000000 | 0.000000 | 0.000000 | 0.0 | 0.0 | 0.0 | 0.000000e+00 | 0.000000 | 0.000000 |
| 4 | 0.000000 | 0.000000 | 0.000000 | 0.0 | 0.0 | 0.000000 | 0.003874 | 0.0 | 0.0 | 0.0 | ... | 0.0 | 0.000000 | 0.000000 | 0.000000 | 0.0 | 0.0 | 0.0 | 0.000000e+00 | 0.000000 | 0.000000 |
| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
| 249 | 0.000000 | 0.000000 | 0.000000 | 0.0 | 0.0 | 0.000000 | 0.000000 | 0.0 | 0.0 | 0.0 | ... | 0.0 | 0.000000 | 0.000000 | 0.000000 | 0.0 | 0.0 | 0.0 | 3.937008e-03 | 0.000000 | 0.000000 |
| 250 | 0.000000 | 0.000000 | 0.000000 | 0.0 | 0.0 | 0.000000 | 0.000000 | 0.0 | 0.0 | 0.0 | ... | 0.0 | 0.000000 | 0.000000 | 0.000000 | 0.0 | 0.0 | 0.0 | 6.299213e-05 | 0.003874 | 0.000000 |
| 251 | 0.000000 | 0.000000 | 0.000000 | 0.0 | 0.0 | 0.000000 | 0.000000 | 0.0 | 0.0 | 0.0 | ... | 0.0 | 0.000000 | 0.000126 | 0.003811 | 0.0 | 0.0 | 0.0 | 3.035766e-17 | 0.000000 | 0.000000 |
| 252 | 0.000000 | 0.000000 | 0.000000 | 0.0 | 0.0 | 0.000000 | 0.000000 | 0.0 | 0.0 | 0.0 | ... | 0.0 | 0.003748 | 0.000000 | 0.000189 | 0.0 | 0.0 | 0.0 | 0.000000e+00 | 0.000000 | 0.000000 |
| 253 | 0.000000 | 0.000000 | 0.000000 | 0.0 | 0.0 | 0.000000 | 0.000000 | 0.0 | 0.0 | 0.0 | ... | 0.0 | 0.000000 | 0.000000 | 0.000000 | 0.0 | 0.0 | 0.0 | 0.000000e+00 | 0.000000 | 0.003937 |
254 rows × 250 columns
Sequential pairwise slice alignment plots
pis = [pi12, pi23, pi34] slices = [slice1, slice2, slice3, slice4] new_slices, _, _ = stack_slices_pairwise(slices, pis)
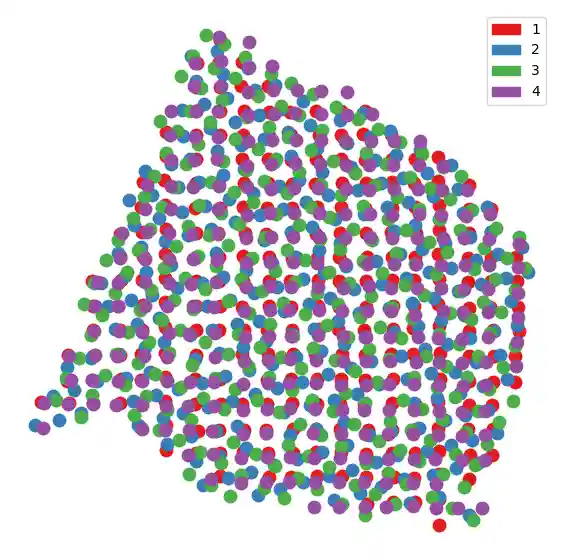
Now that we’ve aligned the spatial coordinates, we can plot them all on the same coordinate system.
slice_colors = ["#e41a1c", "#377eb8", "#4daf4a", "#984ea3"] plt.figure(figsize=(7, 7)) for i in range(len(new_slices)): plot_slice(new_slices[i], slice_colors[i], s=400) plt.legend( handles=[ mpatches.Patch(color=slice_colors[0], label="1"), mpatches.Patch(color=slice_colors[1], label="2"), mpatches.Patch(color=slice_colors[2], label="3"), mpatches.Patch(color=slice_colors[3], label="4"), ] ) plt.gca().invert_yaxis() plt.axis("off") plt.show()
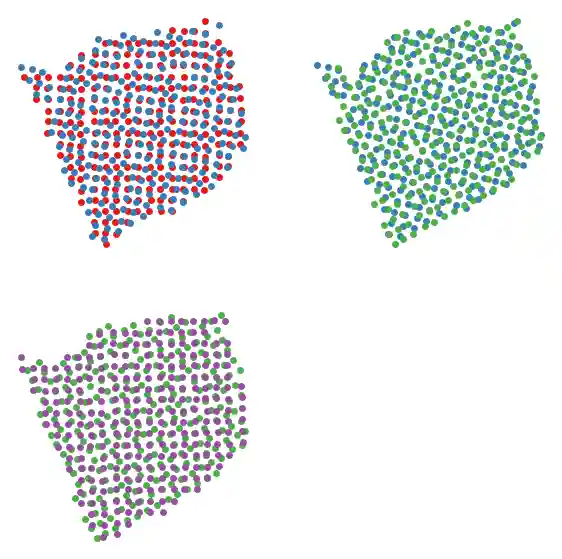
We can also plot pairwise layers together.
slice_colors = ["#e41a1c", "#377eb8", "#4daf4a", "#984ea3"] fig, axs = plt.subplots(2, 2, figsize=(7, 7)) plot_slice(new_slices[0], slice_colors[0], ax=axs[0, 0]) plot_slice(new_slices[1], slice_colors[1], ax=axs[0, 0]) plot_slice(new_slices[1], slice_colors[1], ax=axs[0, 1]) plot_slice(new_slices[2], slice_colors[2], ax=axs[0, 1]) plot_slice(new_slices[2], slice_colors[2], ax=axs[1, 0]) plot_slice(new_slices[3], slice_colors[3], ax=axs[1, 0]) fig.delaxes(axs[1, 1]) plt.show()
We can also plot the slices in 3-D.
import plotly.express as px import plotly.io as pio pio.renderers.default = "notebook" slices_colors = ["#e41a1c", "#377eb8", "#4daf4a", "#984ea3"] # scale the distance between layers z_scale = 2 values = [] for i, L in enumerate(new_slices): for x, y in L.obsm["spatial"]: values.append([x, y, i * z_scale, str(i)]) df = pd.DataFrame(values, columns=["x", "y", "z", "slice"]) fig = px.scatter_3d( df, x="x", y="y", z="z", color="slice", color_discrete_sequence=slice_colors ) fig.update_layout(scene_aspectmode="data") fig.show()
Center Alignment
First, we will read in and preprocess the data (if you ran pairwise_align above, it will be altered).
slices = load_slices(data_dir, ["slice1", "slice2", "slice3", "slice4"]) slice1, slice2, slice3, slice4 = slices
Run PASTE center_align.
slices = [slice1, slice2, slice3, slice4] initial_slice = slice1.copy() lmbda = len(slices) * [1 / len(slices)]
Now, for center alignment, we can provide initial mappings between the center and original slices to PASTE to improve the algorithm. However, note this is optional.
slices, _ = get_common_genes(slices) b = [] for i in range(len(slices)): b.append( torch.Tensor( match_spots_using_spatial_heuristic(slices[0].X, slices[i].X) ).double() )
start = time.time() center_slice, pis = center_align( initial_slice, slices, lmbda, random_seed=5, pi_inits=b ) print("Runtime: " + str(time.time() - start))
(INFO) (paste.py) (10-Jan-25 17:16:56) GPU is not available, resorting to torch CPU. (INFO) (paste.py) (10-Jan-25 17:16:56) Solving Center Mapping NMF Problem. (INFO) (paste.py) (10-Jan-25 17:16:58) Iteration: 0 (INFO) (paste.py) (10-Jan-25 17:16:58) Solving Pairwise Slice Alignment Problem. (INFO) (paste.py) (10-Jan-25 17:16:58) Slice 0 (INFO) (paste.py) (10-Jan-25 17:16:59) Slice 1
/opt/hostedtoolcache/Python/3.12.8/x64/lib/python3.12/site-packages/sklearn/decomposition/_nmf.py:1742: ConvergenceWarning: Maximum number of iterations 200 reached. Increase it to improve convergence.
(INFO) (paste.py) (10-Jan-25 17:16:59) Slice 2 (INFO) (paste.py) (10-Jan-25 17:16:59) Slice 3 (INFO) (paste.py) (10-Jan-25 17:16:59) center_ot done (INFO) (paste.py) (10-Jan-25 17:16:59) Solving Center Mapping NMF Problem. (INFO) (paste.py) (10-Jan-25 17:17:02) Objective -13.865903767431423 | Difference: 13.865903767431423 (INFO) (paste.py) (10-Jan-25 17:17:02) Iteration: 1 (INFO) (paste.py) (10-Jan-25 17:17:02) Solving Pairwise Slice Alignment Problem. (INFO) (paste.py) (10-Jan-25 17:17:02) Slice 0 (INFO) (paste.py) (10-Jan-25 17:17:02) Slice 1
/opt/hostedtoolcache/Python/3.12.8/x64/lib/python3.12/site-packages/sklearn/decomposition/_nmf.py:1742: ConvergenceWarning: Maximum number of iterations 200 reached. Increase it to improve convergence.
(INFO) (paste.py) (10-Jan-25 17:17:02) Slice 2 (INFO) (paste.py) (10-Jan-25 17:17:02) Slice 3 (INFO) (paste.py) (10-Jan-25 17:17:02) center_ot done (INFO) (paste.py) (10-Jan-25 17:17:02) Solving Center Mapping NMF Problem. (INFO) (paste.py) (10-Jan-25 17:17:05) Objective 1.3829621916807069 | Difference: 15.24886595911213 (INFO) (paste.py) (10-Jan-25 17:17:05) Iteration: 2 (INFO) (paste.py) (10-Jan-25 17:17:05) Solving Pairwise Slice Alignment Problem. (INFO) (paste.py) (10-Jan-25 17:17:05) Slice 0 (INFO) (paste.py) (10-Jan-25 17:17:05) Slice 1
/opt/hostedtoolcache/Python/3.12.8/x64/lib/python3.12/site-packages/sklearn/decomposition/_nmf.py:1742: ConvergenceWarning: Maximum number of iterations 200 reached. Increase it to improve convergence.
(INFO) (paste.py) (10-Jan-25 17:17:05) Slice 2 (INFO) (paste.py) (10-Jan-25 17:17:05) Slice 3 (INFO) (paste.py) (10-Jan-25 17:17:05) center_ot done (INFO) (paste.py) (10-Jan-25 17:17:05) Solving Center Mapping NMF Problem. (INFO) (paste.py) (10-Jan-25 17:17:08) Objective 1.3880932065404366 | Difference: 0.005131014859729666 (INFO) (paste.py) (10-Jan-25 17:17:08) Iteration: 3 (INFO) (paste.py) (10-Jan-25 17:17:08) Solving Pairwise Slice Alignment Problem. (INFO) (paste.py) (10-Jan-25 17:17:08) Slice 0 (INFO) (paste.py) (10-Jan-25 17:17:08) Slice 1
/opt/hostedtoolcache/Python/3.12.8/x64/lib/python3.12/site-packages/sklearn/decomposition/_nmf.py:1742: ConvergenceWarning: Maximum number of iterations 200 reached. Increase it to improve convergence.
(INFO) (paste.py) (10-Jan-25 17:17:08) Slice 2 (INFO) (paste.py) (10-Jan-25 17:17:08) Slice 3 (INFO) (paste.py) (10-Jan-25 17:17:08) center_ot done (INFO) (paste.py) (10-Jan-25 17:17:08) Solving Center Mapping NMF Problem. (INFO) (paste.py) (10-Jan-25 17:17:11) Objective 1.3915232061917202 | Difference: 0.003429999651283655 (INFO) (paste.py) (10-Jan-25 17:17:11) Iteration: 4 (INFO) (paste.py) (10-Jan-25 17:17:11) Solving Pairwise Slice Alignment Problem. (INFO) (paste.py) (10-Jan-25 17:17:11) Slice 0 (INFO) (paste.py) (10-Jan-25 17:17:11) Slice 1
/opt/hostedtoolcache/Python/3.12.8/x64/lib/python3.12/site-packages/sklearn/decomposition/_nmf.py:1742: ConvergenceWarning: Maximum number of iterations 200 reached. Increase it to improve convergence.
(INFO) (paste.py) (10-Jan-25 17:17:11) Slice 2 (INFO) (paste.py) (10-Jan-25 17:17:11) Slice 3 (INFO) (paste.py) (10-Jan-25 17:17:11) center_ot done (INFO) (paste.py) (10-Jan-25 17:17:11) Solving Center Mapping NMF Problem. (INFO) (paste.py) (10-Jan-25 17:17:14) Objective 1.3930712361281365 | Difference: 0.0015480299364163397 (INFO) (paste.py) (10-Jan-25 17:17:14) Iteration: 5 (INFO) (paste.py) (10-Jan-25 17:17:14) Solving Pairwise Slice Alignment Problem. (INFO) (paste.py) (10-Jan-25 17:17:14) Slice 0 (INFO) (paste.py) (10-Jan-25 17:17:14) Slice 1
/opt/hostedtoolcache/Python/3.12.8/x64/lib/python3.12/site-packages/sklearn/decomposition/_nmf.py:1742: ConvergenceWarning: Maximum number of iterations 200 reached. Increase it to improve convergence.
(INFO) (paste.py) (10-Jan-25 17:17:14) Slice 2 (INFO) (paste.py) (10-Jan-25 17:17:14) Slice 3 (INFO) (paste.py) (10-Jan-25 17:17:14) center_ot done (INFO) (paste.py) (10-Jan-25 17:17:14) Solving Center Mapping NMF Problem. (INFO) (paste.py) (10-Jan-25 17:17:17) Objective 1.395250483498415 | Difference: 0.0021792473702784143 (INFO) (paste.py) (10-Jan-25 17:17:17) Iteration: 6 (INFO) (paste.py) (10-Jan-25 17:17:17) Solving Pairwise Slice Alignment Problem. (INFO) (paste.py) (10-Jan-25 17:17:17) Slice 0 (INFO) (paste.py) (10-Jan-25 17:17:17) Slice 1
/opt/hostedtoolcache/Python/3.12.8/x64/lib/python3.12/site-packages/sklearn/decomposition/_nmf.py:1742: ConvergenceWarning: Maximum number of iterations 200 reached. Increase it to improve convergence.
(INFO) (paste.py) (10-Jan-25 17:17:17) Slice 2 (INFO) (paste.py) (10-Jan-25 17:17:17) Slice 3 (INFO) (paste.py) (10-Jan-25 17:17:18) center_ot done (INFO) (paste.py) (10-Jan-25 17:17:18) Solving Center Mapping NMF Problem. (INFO) (paste.py) (10-Jan-25 17:17:20) Objective 1.3953790526176948 | Difference: 0.00012856911927983106 (INFO) (paste.py) (10-Jan-25 17:17:21) Center slice computed. Runtime: 24.87651538848877
/opt/hostedtoolcache/Python/3.12.8/x64/lib/python3.12/site-packages/sklearn/decomposition/_nmf.py:1742: ConvergenceWarning: Maximum number of iterations 200 reached. Increase it to improve convergence.
Again, we can run center align without providing intial mappings below.
# center_slice, pis = paste.center_align(initial_slice, slices, lmbda, random_seed = 5)
center_slice returns an AnnData object that also includes the low dimensional representation of our inferred center slice.
center_slice.uns["paste_W"]
array([[2.56406116e-02, 1.17735608e+00, 1.45992053e-01, ...,
1.67005408e-02, 8.65816920e-03, 1.64249334e-02],
[1.93240013e-03, 3.38372966e-01, 1.80787149e-01, ...,
1.61882753e-02, 1.01921081e-03, 4.46527714e-02],
[4.19665832e-01, 1.88694801e-01, 2.90303240e-02, ...,
4.75023311e-02, 1.62389887e-02, 1.17923350e-01],
...,
[2.91596165e-02, 8.19205072e-03, 1.07181455e-01, ...,
4.61294426e-04, 4.20524981e-04, 2.33305738e-01],
[4.31159630e-02, 3.82986839e-02, 1.78688007e-01, ...,
9.82425693e-03, 3.23265860e-03, 2.58806995e-01],
[1.83314337e-01, 1.68948804e-05, 2.22536788e-02, ...,
1.16837119e-05, 4.58040466e-03, 9.97049918e-02]])
center_slice.uns["paste_H"]
array([[9.00595555e-01, 1.28095211e-01, 7.99820106e-02, ...,
5.45672185e-03, 4.42535682e-02, 2.77099853e-02],
[1.84556880e+00, 1.12907521e-01, 3.19053463e-01, ...,
1.01912357e-01, 1.03440813e-01, 3.08299135e-02],
[1.93982105e+00, 1.42403630e-01, 1.30713051e-01, ...,
7.35120019e-02, 3.50568645e-02, 2.86018512e-02],
...,
[3.05294205e+00, 1.88580791e-01, 3.20976581e-01, ...,
2.51081658e-02, 5.00736543e-06, 1.10129713e-02],
[9.20159500e-01, 1.65184151e-01, 1.04867309e-01, ...,
1.26153742e-02, 1.25054026e-02, 2.44712742e-02],
[5.73756503e+00, 7.43537812e-04, 3.13147999e-01, ...,
6.80617759e-06, 3.99149863e-02, 2.55629904e-13]])
Center slice alignment plots
Next, we can use the outputs of center_align to align the slices.
center, new_slices, _, _ = stack_slices_center(center_slice, slices, pis)
Now that we’ve aligned the spatial coordinates, we can plot them all on the same coordinate system. Note the center slice is not plotted.
center_color = "orange" slices_colors = ["#e41a1c", "#377eb8", "#4daf4a", "#984ea3"] plt.figure(figsize=(7, 7)) plot_slice(center, center_color, s=400) for i in range(len(new_slices)): plot_slice(new_slices[i], slices_colors[i], s=400) plt.legend( handles=[ mpatches.Patch(color=slices_colors[0], label="1"), mpatches.Patch(color=slices_colors[1], label="2"), mpatches.Patch(color=slices_colors[2], label="3"), mpatches.Patch(color=slices_colors[3], label="4"), ] ) plt.gca().invert_yaxis() plt.axis("off") plt.show()
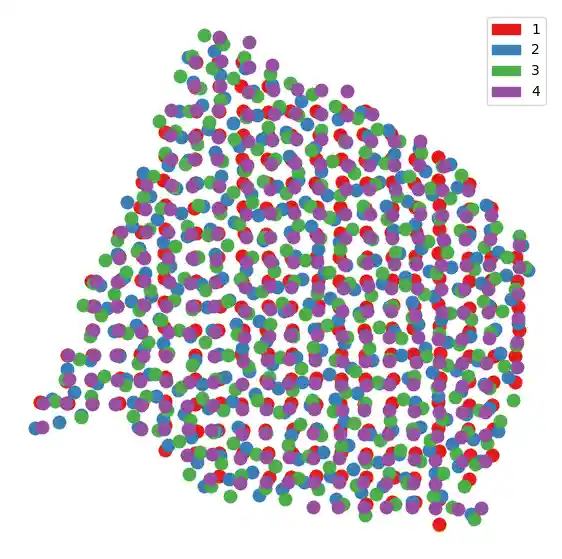
Next, we plot each slice compared to the center.
Note that since we used slice1 as the coordinates for the center slice, they remain the same, and thus we cannot see both in our plots below.
center_color = "orange" slice_colors = ["#e41a1c", "#377eb8", "#4daf4a", "#984ea3"] fig, axs = plt.subplots(2, 2, figsize=(7, 7)) plot_slice(center, center_color, ax=axs[0, 0]) plot_slice(new_slices[0], slice_colors[0], ax=axs[0, 0]) plot_slice(center, center_color, ax=axs[0, 1]) plot_slice(new_slices[1], slice_colors[1], ax=axs[0, 1]) plot_slice(center, center_color, ax=axs[1, 0]) plot_slice(new_slices[2], slice_colors[2], ax=axs[1, 0]) plot_slice(center, center_color, ax=axs[1, 1]) plot_slice(new_slices[3], slice_colors[3], ax=axs[1, 1]) plt.show()