Snakemake
Readability and automation
With Snakemake, data analysis workflows are defined via an easy to read, adaptable, yet powerful specification language on top of Python. Steps are defined by "rules", which denote how to generate a set of output files from a set of input files (e.g. using a shell command). Wildcards (in curly braces) provide generalization. Dependencies between rules are determined automatically.
rule select_by_country:
input:
"data/worldcitiespop.csv"
output:
"by-country/{country}.csv"
shell:
"xsv search -s Country '{wildcards.country}' "
"{input} > {output}"
Portability
By integration with the Conda package manager and containers, all software dependencies of each workflow step are automatically deployed upon execution.
rule select_by_country:
input:
"data/worldcitiespop.csv"
output:
"by-country/{country}.csv"
conda:
"envs/xsv.yaml"
shell:
"xsv search -s Country '{wildcards.country}' "
"{input} > {output}"
Scripting integration
Rapidly implement analysis steps via direct script and jupyter notebook integration supporting Python, R, Julia, Rust, Bash, without requiring any boilerplate code.
rule select_by_country:
input:
"data/worldcitiespop.csv"
output:
"by-country/{country}.csv"
script:
"scripts/select_by_country.R"
rule convert_to_pdf:
input:
"{prefix}.svg"
output:
"{prefix}.pdf"
wrapper:
"0.47.0/utils/cairosvg"
"Turing completeness"
Being a syntactical extension of Python, you can implement arbitrary logic beyond the plain definition of rules. Rules can be generated conditionally, arbitrary Python logic can be used to perform aggregations, configuration and metadata can be obtained and postprocessed in any required way.
def get_data(wildcards):
# use arbitrary Python logic to
# aggregate over the required input files
return ...
rule plot_histogram:
input:
get_data
output:
"plots/hist.svg"
script:
"scripts/plot-hist.py"
Human Readability
The logic of production workflows can become complex by involving lots of lookups and dynamic decisions. Snakemake offers semantic helper functions for lookups, branching and aggregation that avoid the need for plain Python code as shown above, and allow to express complex logic in a human-readable and self-contained way.
rule plot_histogram:
input:
branch(
lookup(dpath="histogram/somedata", within=config),
then="data/somedata.txt",
otherwise="data/someotherdata.txt"
)
output:
"plots/hist.svg"
script:
"scripts/plot-hist.py"
Dynamic workflows
Snakemake allows to define workflows that are dynamically updated at runtime. By defining so-called checkpoints, the workflow can be dynamically adapted at runtime. Further, input can be provided as Python queues, thereby enabling a workflow to continuously receive new input data (e.g. while a certain measurement is conducted).
rule all:
input:
from_queue(all_results, finish_sentinel=...)
checkpoint somestep:
input:
"samples/{sample}.txt"
output:
"somestep/{sample}.txt"
shell:
"somecommand {input} > {output}"
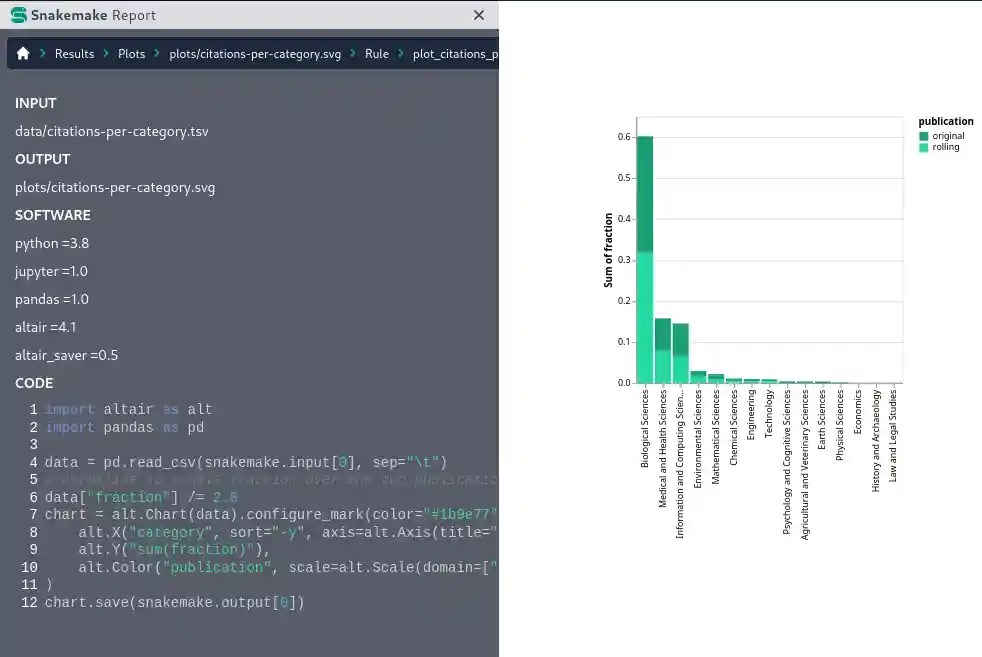
Transparency and data provenance
Automatic, interactive, self-contained reports ensure full transparency from results down to used steps, parameters, code, and software. The reports can moreover contain embedded results (from images, to PDFs and even interactive HTML) enabling a comprehensive reporting that combines analysis results with data provenance information.
Scalability
Workflows scale seamlessly from single to multicore, clusters or the cloud, without modification of the workflow definition and automatic avoidance of redundant computations.
Configurability
Snakemake is extremely flexible and configurable. Numerous options allow adapt the behavior to the needs of the data analysis at hand and the underlying infrastructure. Options can be provided via the command line interface or persisted via system-wide, user-specific, and workflow specific profiles.
executor: slurm
software-deployment-method:
- conda
latency-wait: 60
default-storage-provider: fs
shared-fs-usage:
- persistence
- software-deployment
- sources
- source-cache
local-storage-prefix:
/local/work/$USER/snakemake-scratch
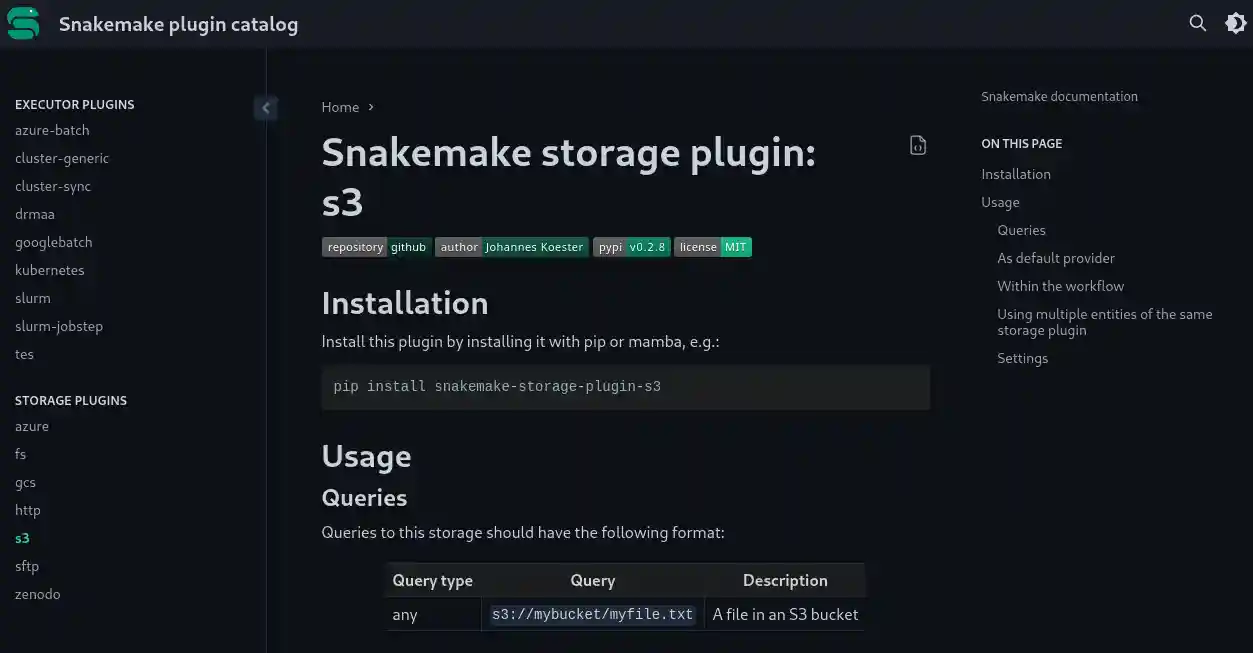
Extensibility
Snakemake has a powerful plugin system that allows to extend various functionalities with alternative implementations. Via stable and well-defined interfaces, plugins can evolve independently of Snakemake, and mutual update requirements are minimized. Currently, execution backends and remote storage support is implemented via plugins. In the future, we will extend this to other areas, such as workflow scheduling, reporting, software deployment, and more.
Authors and Contributors ⓘ
- Johannes Köster
- Per Unneberg
- Chris Tomkins-Tinch
- Rasmus Ågren
- Tim Booth
- Vanessasaurus
- snakemake-bot
- Filipe G. Vieira
- Henning Timm
- David Laehnemann
- Chris Burr
- Manuel Holtgrewe
- Marcel Martin
- Michael Hall
- Cade Mirchandani
- Ryan C. Thompson
- Wibowo Arindrarto
- Elmar Pruesse
- Morten Enemark Lund
- Felix Mölder
- Aoran Hu
- Ryan Dale
- Ben Beasley
- Fredrik Boulund
- Peter Van Dyken
- Simon Ye
- Derek Croote
- Oliver Stolpe
- Jake VanCampen
- Justin Fear
- Kemal Eren
- Mattias de Hollander
- Maarten-vd-Sande
- Dr. K. D. Murray
- Michael K. Wilkinson
- Maarten Kooyman
- Kim
- Christian Arnold
- Jermiah Joseph
- Jay Hesselberth
- Hyeshik Chang
- Cornelius Roemer
- Troy Comi
- John Eppley
- Mattias Frånberg
- Felix Wiegand
- Christian Meesters
- mhulsman
- Nils Homer
- Patrik Smeds
- Kyle Beauchamp
- Liang-Bo Wang
- Joseph K Aicher
- Mike Taves
- Lars Bilke
- Kevin Sayers
- C. Titus Brown
- Anfeng Li
- Sven Twardziok
- Sultan Orazbayev
- Matt Shirley
- G. D. McBain
- endrebak
- Adam Labadorf
- Anthony Underwood
- Elias Kuthe
- Haizi Zheng
- Jan Forster
- John Blischak
- Don Freed
- Dmitry Kalinkin
- Marco-Masera
- Peter Cock
- Vito Zanotelli
- Shinji Matsumoto
- Rohan-Ibn-Tariq
- Benjamin Yeh
- Ryan A. Hagenson
- Sichong
- Silas Kieser
- Vlad Savelyev
- Soo Lee
- Sven Schrinner
- wligtenberg
- Hatem
- Florian R. Hölzlwimmer
- Thomas Vandal
- Tomás Di Domenico
- rebecca-palmer
- darrin t schultz
- Bruno P. Kinoshita
- Ashwin V. Mohanan
- Andreas Wilm
- crimsonDaMi
- Arya Massarat
- Christian Brueffer
- Christopher Schröder
- David Alexander
- David Koppstein
- Jeremy Leipzig
- Joona Lehtomäki
- Koen van Greevenbroek
- Kyle Meyer
- Nicolas Ochsner
- Nils Giordano
- Pay Giesselmann
- Yoshiki Vázquez Baeza
- Alex Leonard
- Sebastian Schmidt
- nikostr
- jlncrnt
- dkuzminov
- cclienti
- Zeb Burke-Conte
- Vince
- Till Hartmann
- Hielke Walinga
- Wolfgang Kopp
- Ward D
- DrYak
- Ethan Holleman
- Sebastian
- Jesse Connell
- John Marshall
- Jon Stutters
- Patrick Kunzmann
- Norbert Auer
- Lucas Frérot
- Michael R. Crusoe
- Matthew Monk
- Hugo Lapré
- Doğukan
- J.J.
- Ezra Herman
- JS Légaré
- Heath O'Brien
- Dmytro Kazanzhy
- Devon Ryan
- Christian
- Foivos Gypas
- Alexander Kleinjohann
- Frédéric Chevalier
- Thomas Weber
- Thom Griffioen
- Roy Jacobson
- Rich Abdill
- Peter Schiffels
- Nelis Drost
- Naveen
- Michael Schubert
- Marawan Abdelgawad
- Lance Parsons
- Jo Hausmann
- Adam Morris
- Jens Zentgraf
- Sebastian Ohlmann
- Sean Davis
- Seth Ariel Green
- Quinn Blenkinsop
- Raphael Müller
- ScottMastro
- Renan Valieris
- Renato Alves
- RezaMadi
- Samuel Gaist
- Rick Tankard
- Rob Schaefer
- Sam-Tygier
- Robben Migacz
- Sam Nicholls
- Robert Schauner
- Ryunosuke O'Neil
- Romain Feron
- Rodrigo Luger
- Connor Jops
- Matt Stone
- Matthias Peter
- Matthias Wolf
- Michal Stolarczyk
- Mike DePalatis
- Mitchell Robert Vollger
- Mohammad Samman
- Moustapha Sall
- Murillo F. Rodrigues
- Nicholas A. Del Grosso
- Nick Semenkovich
- Noah
- Oliver Küchler
- Paul Bransford
- Paul K. Korir
- Paul L. Maurizio
- Paul Moore
- Perry
- Pete Bachant
- Peter Steinbach
- Philipp Helo Rehs
- Pierre Marijon
- Zbigniew Jędrzejewski-Szmek
- Bailey Harrington
- bcouturi
- Bilgehan Nevruz
- bingxiao
- chaen
- Eric Jelli
- Johannes HAMPP
- johannaschmitz
- maxim-h
- moschetti
- nbelakovski
- nhartwic
- ningOTI
- ocaisa
- olgert denas
- phmoferring
- rlehnigk
- roshnipatel
- snajder-r
- stydodt
- tdayris
- Valentin Schneider-Lunitz
- xhejtman
- Žiga Avsec
- Stefan Pfenninger-Lee
- Stephan Druskat
- Susana Posada Céspedes
- Tadas Bareikis
- Tanguy Lallemand
- Taylor Reiter
- Terry Jones
- Tet Woo Lee
- Thomas Grainger
- Thomas Mulvaney
- Thomas Sibley
- Tim Heap
- Tim Tröndle
- Tobias Ternent
- Travis Wrightsman
- Valentin Pestel
- Valentyn Bezshapkin
- Vincent
- Vini Salazar
- pirovc
- Vladimir Mikryukov
- Wim Jeantine-Glenn
- Yaman Qalieh
- Yaroslav Halchenko
- Ye Yuan
- CowanCS1
- Daniel Lusk
- Dario Beraldi
- Derrick Miller
- Diego Rubert
- Dr. Fabian Schlegel
- Dustin Rodrigues
- Edmund Miller
- Edouard Choinière
- Egor Kosaretskiy
- Emmanouil "Manolis" Maragkakis
- Eric Normandeau
- Fabian Neumann
- Flo
- Fong Chun Chan
- Afonso Santos
- Frank Löffler
- Frankie Robertson
- Fritjof Lammers
- Jean-Sebastien Gounot
- Gaspard Reulet
- George Wu
- Giacomo Tagliabue
- gibran hemani
- Adam Ciuris
- Alistair Miles
- Andrew Berger
- Andrii Oriekhov
- Aniket Pradhan
- arnikz
- BAKEZQ
- Balázs Brankovics
- Ben Evans
- Ben Jolly
- Branch Vincent
- Brian Fulton-Howard
- Brice Letcher
- Byron J. Smith
- Cail McLean Daley
- Carl Mathias Kobel
- Chang Ye
- Chaz Reid
- Charlie Pauvert
- Chris Lamb
- Christian Klarhorst
- ChristofferCOASD
- Clemens Lange
- ClementFombonne
- Colin J. Brislawn
- Justin Hiemstra
- Karatuğ Ozan Bircan
- Karel Břinda
- Kevin Hoffschlag
- Knut Dagestad Rand
- Kostis Anagnostopoulos
- Kyle Johnsen
- LarsStegemanGT
- Leonardo Schwarz
- Leopoldo Pla Sempere
- Louis Vignoli
- Ludwig Neste
- Luis Kress
- Lukas Klein
- LvKvA
- Maarten0110
- Maciek Bąk
- Marcel Bargull
- Marcin Magnus
- Marco Vidal García
- Mark Keller
- Martin Holub
- Martin Stancsics
- Mathieu Bernard
- Matthew Feickert
- Giles Harper-Donnelly
- Giulio Centorame
- Grégoire Denay
- Hamdiye Uzuner
- Henry Webel
- Ido Tamir
- Jakub Kaczmarzyk
- James Chuang
- James Shaw
- Jan Eil
- Jason Greenbaum
- scholtalbers
- Jeremy Magland
- Jesse Bloom
- Jesse Wallace
- Jigar Patel
- Joe Sapp
- Johannes Heuel
- Johannes Schumann
- John Bates
- John Hennig
- John Huddleston
- (major) john (major)
- Josh Cook
- JulioV
Groups, Institutes, Companies, and Organizations ⓘ
- University of Duisburg-Essen
- Science for Life Laboratory
- Broad Institute of MIT and Harvard
- On Sabbatical
- The GLOBE Institute – University of Copenhagen
- CERN
- CUBI Core Unit Bioinformatics, Berlin Institute of Health
- University of Queensland | UQCCR
- UC Santa Cruz
- Icahn School of Medicine at Mount Sinai
- group.one
- @AnyBody
- 上海交通大学
- Karolinska Institutet
- Pairwise
- Netherlands Institute of Ecology (NIOO-KNAW)
- Solynta
- Gekkonid Scientific
- LHCb
- Oyat Consulting
- Data Science @google
- EMBL
- Princess Margaret Cancer Centre, University Health Network
- University of Colorado School of Medicine
- Seoul National University
- @neherlab @nextstrain
- Princeton University
- Spotify
- Universität Duisburg-Essen
- University of Mainz
- @fulcrumgenomics
- Clinical Genomics Uppsala / Uppsala University
- Earth Sciences New Zealand (formerly GNS Science)
- Helmholtz Centre for Environmental Research
- AWS
- University of California, Davis
- BIH
- @cid-harvard
- @novartis
- NTNU
- Regeneron Pharmaceuticals
- Sentieon
- UZH Zurich
- Washington University School of Medicine
- @seqera.io
- Exact Sciences
- University Medicine Essen
- Technical University of Munich
- Université de Montréal
- @cnio-bu
- University of Vienna
- Barcelona Supercomputing Center
- @ENCCS
- ImmunoScape
- Gymrek Lab, UCSD
- @insilicoconsulting
- University of Duisburg Essen
- DKTK/DKFZ
- @TileDB-Inc
- Stanford University
- ETH
- DKRZ
- @BiomeSense
- ETHZ
- University of Helsinki
- Institute for Health Metrics and Evaluation
- @bihealth
- Max Delbrück center for molecular medicine
- @sib-swiss
- Alva Genomics
- University of Pennsylvania
- University College London
- VantAI
- IMS Nanofabrication GmbH
- Sorbonne Université, Paris
- @common-workflow-language
- Brabant Water
- Esox Biologics
- @GenomicsUA @lyft
- Genedata AG
- Bioinformatics Software Engineer at Novartis Institutes for BioMedical Research (NIBR)
- @RWTH-EBC
- @txbiomed
- Data Science Centre, EMBL
- @Syngenta
- University of Chicago
- Center for eResearch, University of Auckland
- @TRON-Bioinformatics
- Saarland University
- University of Colorado Anschutz School of Medicine
- Kahnemnan-Treisman Center
- A. C. Camargo Cancer Center
- @idiap
- WEHI
- University of Utah Center for High Performance Computing
- @nanoporetech
- Anthropic
- @fulcrumgenomics
- Ascend Analytics
- @Elembio
- Medical College of Wisconsin
- University of Pittsburgh / Center for Craniofacial and Dental Genetics
- Freie Universität Berlin
- @USF-HII
- DataDotOrg
- NIAID
- Caltech
- www.hzdr.de
- HHU Düsseldorf
- Seqoia
- Red Hat
- Amsterdam UMC
- @mpinb
- @open-energy-transition
- University of Basel
- Duke-NUS
- @BlueRiverTechnology
- Gustave Roussy
- TU Delft
- German Aerospace Center (DLR)
- IOB
- @open-energy-transition
- Novartis
- LPC Caen - IN2P3 - CNRS
- Sunagawa Lab @ ETH Zürich
- University of Tartu
- Dartmouth College, @dandi, @Debian, @DataLad, @neurodebian, @PyMVPA, @fail2ban
- University of Michigan, Ann Arbor
- WCIP | University of Glasgow
- Helmholtz-Zentrum Dresden-Rossendorf e.V.
- @seqeralabs
- National Institutes of Health
- IBIS (Institut de Biologie Intégrative et des Systèmes)
- TU Berlin
- Roche
- Friedrich Schiller University Jena
- University of Jyväskylä
- Swiss Ornithological Institute
- Montreal Clinical Research Institute
- 4Catalyzer
- @tempuslabs
- Wellcome Sanger Institute
- SPD
- @Adobe
- Westerdijk Fungal Bioidiversity Institute
- Sarepta Therapeutics
- @manaakiwhenua
- Mount Sinai Hospital (@marcoralab)
- CNRS
- The Gladstone Institutes
- Wesleyan University
- Norges Miljø- & Biovitenskapelige Universitet (NMBU)
- Freelancer for hire. Maybe.
- Paul Scherrer Institute
- Morgridge Instute for Research
- 𝐈𝐍𝐑𝐈𝐀🇫🇷 Nat. Inst. for DigitSci & Tech
- Predictive Neuroscience Lab, University Hospital Essen
- Oslo University Hospital
- @JRC-IET, C3
- Georgia Tech
- FGCZ, ETHZ | UZH
- Treelogic & University of Alacant
- Pasqal
- Stockholm Universitetet
- TRON gGmbH Mainz
- Hochschule Darmstadt
- City, University of London
- Harvard, USA
- @Quantco
- LPNHE - CNRS - Sorbonne Université
- University of Wisconsin-Madison
- Institute for Molecular Bioscience, University of Queensland
- @bio-raum @CVUA-RRW
- DTU biosustain
- VBCF
- Stony Brook Medicine
- La Jolla Institute for Allergy and Immunology @LJI-Bioinformatics @IEDB
- Fred Hutchinson Cancer Research Center; Howard Hughes Medical Institute
- CSIRO
- Erlangen Centre for Astroparticle Physics
- @blab @nextstrain
- Daylily Informatics
- Vertex Pharmaceuticals